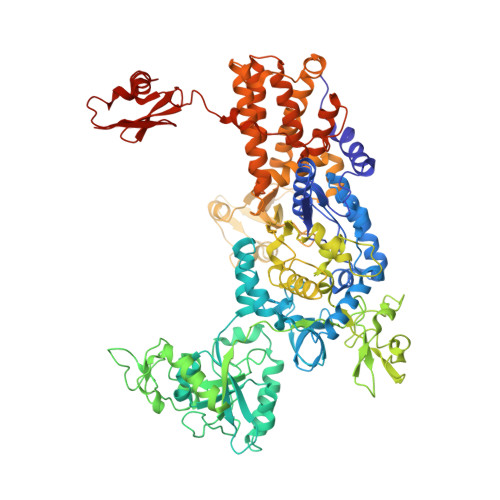
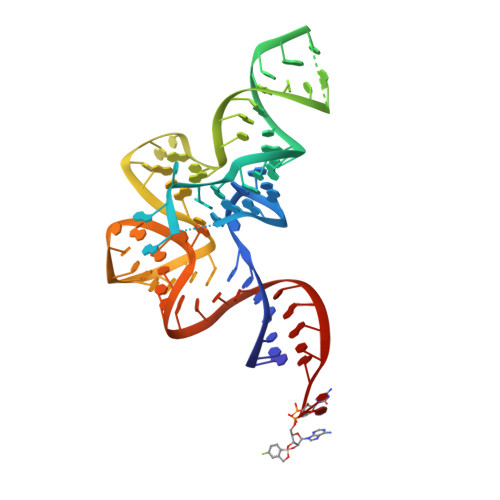
An Antifungal Agent Inhibits an Aminoacyl-tRNA Synthetase by Trapping tRNA in the Editing Site.
Rock, F., Mao, W., Yaremchuk, A., Tukalo, M., Crepin, T., Zhou, H., Zhang, Y., Hernandez, V., Akama, T., Baker, S., Plattner, J., Shapiro, L., Martinis, S.A., Benkovic, S.J., Cusack, S., Alley, M.R.K.(2007) Science 316: 1759
- PubMed: 17588934 Search on PubMed
- DOI: https://doi.org/10.1126/science.1142189
- Primary Citation Related Structures:
2V0C, 2V0G - PubMed Abstract:
Aminoacyl-transfer RNA (tRNA) synthetases, which catalyze the attachment of the correct amino acid to its corresponding tRNA during translation of the genetic code, are proven antimicrobial drug targets. We show that the broad-spectrum antifungal 5-fluoro-1,3-dihydro-1-hydroxy-2,1-benzoxaborole (AN2690), in development for the treatment of onychomycosis, inhibits yeast cytoplasmic leucyl-tRNA synthetase by formation of a stable tRNA(Leu)-AN2690 adduct in the editing site of the enzyme. Adduct formation is mediated through the boron atom of AN2690 and the 2'- and 3'-oxygen atoms of tRNA's3'-terminal adenosine. The trapping of enzyme-bound tRNA(Leu) in the editing site prevents catalytic turnover, thus inhibiting synthesis of leucyl-tRNA(Leu) and consequentially blocking protein synthesis. This result establishes the editing site as a bona fide target for aminoacyl-tRNA synthetase inhibitors.
- Anacor Pharmaceuticals, Incorporated, 1060 East Meadow Circle, Palo Alto, CA 94303, USA.
Organizational Affiliation: