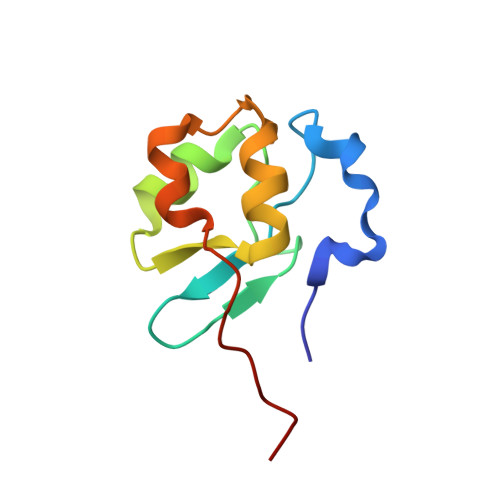
Solution Structure of the Brk Domains from Chd7
Allen, M.D., Religa, T.L., Freund, S.M.V., Bycroft, M.(2007) J Mol Biology 371: 1135
- PubMed: 17603073 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.06.007
- Primary Citation Related Structures:
2V0E, 2V0F - PubMed Abstract:
CHD7 is a member of the chromodomain helicase DNA binding domain (CHD) family of ATP-dependent chromatin remodelling enzymes. It is mutated in CHARGE syndrome, a multiple congenital anomaly condition. CHD7 is one of a subset of CHD proteins, unique to metazoans that contain the BRK domain, a protein module also found in the Brahma/BRG1 family of helicases. We describe here the NMR solution structure of the two BRK domains of CHD7. Each domain has a compact betabetaalphabeta fold. The second domain has a C-terminal extension consisting of two additional helices. The structure differs from those of other domains present in chromatin-associated proteins.
- MRC Centre for Protein Engineering, Hills Road, Cambridge, CB2 2QH, UK.
Organizational Affiliation: