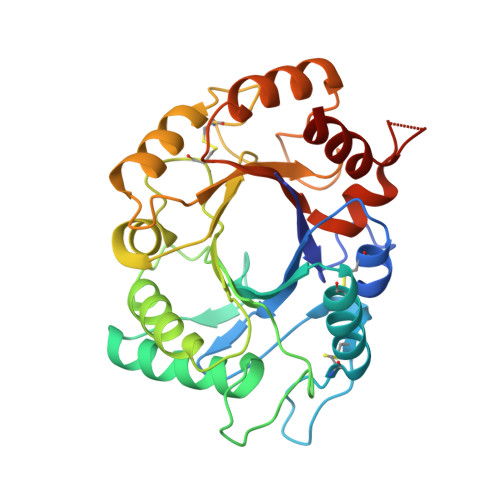
Structure of Saccharomyces Cerevisiae Chitinase 1 and Screening-Based Discovery of Potent Inhibitors.
Hurtado-Guerrero, R., Van Aalten, D.M.F.(2007) Chem Biol 14: 589
- PubMed: 17524989 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2007.03.015
- Primary Citation Related Structures:
2UY2, 2UY3, 2UY4, 2UY5 - PubMed Abstract:
Chitinases hydrolyse the beta(1,4)-glycosidic bonds of chitin, an essential fungal cell wall component. Genetic data on a subclass of fungal family 18 chitinases have suggested a role in cell wall morphology. Specific inhibitors of these enzymes would be useful as tools to study their role in cell wall morphogenesis and could possess antifungal properties. Here, we describe the crystallographic structure of a fungal "plant-type" family 18 chitinase, that of Saccharomyces cerevisiae CTS1. The enzyme is active against 4-methylumbelliferyl chitooligosaccharides and displays an unusually low pH optimum for activity. A library screen against ScCTS1 yielded hits with Ki 's as low as 3.2 microM. Crystal structures of ScCTS1 in complex with inhibitors from three series reveal striking mimicry of carbohydrate substrate by small aromatic moieties and a pocket that could be further exploited in optimization of these inhibitors.
- Division of Biological Chemistry & Molecular Microbiology, School of Life Sciences, University of Dundee, Dundee DD1 5EH, Scotland, UK.
Organizational Affiliation: