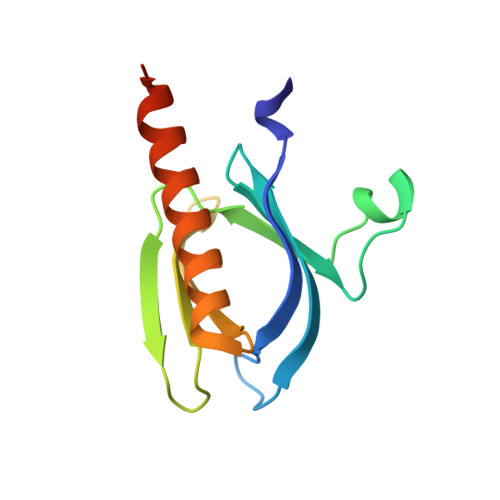
Novel Inositol Phospholipid Headgroup Surrogate Crystallised in the Pleckstrin Homology Domain of Protein Kinase Balpha.
Mills, S.J., Komander, D., Trusselle, M.N., Safrany, S.T., Van Aalten, D.M.F., Potter, B.V.L.(2007) ACS Chem Biol 2: 242
- PubMed: 17432822 Search on PubMed
- DOI: https://doi.org/10.1021/cb700019r
- Primary Citation Related Structures:
2UVM - PubMed Abstract:
Protein kinase B (PKB/Akt) plays a key role in cell signaling. The PH domain of PKB binds phosphatidylinositol 3,4,5-trisphosphate translocating PKB to the plasma membrane for activation by 3-phosphoinositide-dependent protein kinase 1. The crystal structure of the headgroup inositol 1,3,4,5-tetrakisphosphate Ins(1,3,4,5)P4-PKB complex facilitates in silico ligand design. The novel achiral analogue benzene 1,2,3,4-tetrakisphosphate (Bz(1,2,3,4)P4) possesses phosphate regiochemistry different from that of Ins(1,3,4,5)P4 and surprisingly binds with similar affinity as the natural headgroup. Bz(1,2,3,4)P4 co-crystallizes with the PKBalpha PH domain in a fashion also predictable in silico. The 2-phosphate of Bz(1,2,3,4)P4 does not interact with any residue, and the D5-phosphate of Ins(1,3,4,5)P4 is not mimicked by Bz(1,2,3,4)P4. Bz(1,2,3,4)P4 is an example of a simple inositol phosphate surrogate crystallized in a protein, and this approach could be applied to design modulators of inositol polyphosphate binding proteins.