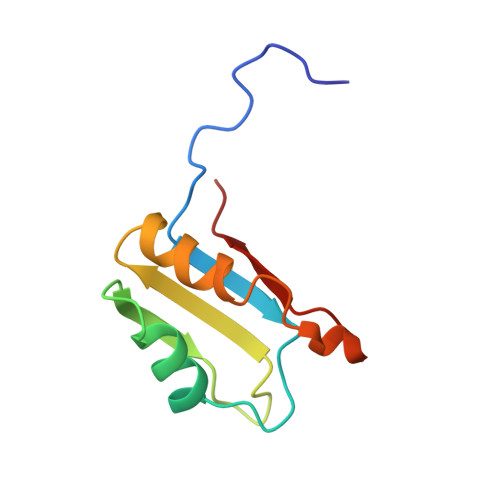
Tertiary structure of RBD2 and backbone dynamics of RBD1 and RBD2 of the human U1A protein determined by NMR spectroscopy.
Lu, J., Hall, K.B.(1997) Biochemistry 36: 10393-10405
- PubMed: 9265619 Search on PubMed
- DOI: https://doi.org/10.1021/bi9709811
- Primary Citation Related Structures:
2U1A - PubMed Abstract:
The human U1A protein has two putative RNA binding domains, one at the N-terminal region of the protein (RBD1) and the other at the C-terminal end (RBD2). RBD1 binds tightly and specifically to one of the stem loops of the U1 snRNA, as well as to its own 3'-UTR. In contrast, RBD2 does not appear to associate with any RNA. The two domains share 25% amino acid identity, and both have the same betaalphabeta-betaalphabeta secondary structure fold. In this work, 13C/15N/1H multidimensional NMR methods were used to obtain side-chain assignments for RBD2, and then the tertiary structure was calculated using a distance geometry/simulated annealing algorithm that employs pairwise Gaussian metrization. RBD2 is shown to fold into an alpha/beta sandwich with a four-stranded antiparallel beta-sheet, which is the typical global topology of these domains. Specific structural features of RBD2 include a beta-bulge in beta2, N-capping boxes for both alpha-helices, and an extremely shallow twist of its beta-sheet. The 15N backbone dynamics of these two structurally homologous RBDs are significantly different, compared using order parameters and T2 exchange terms in the Lipari and Szabo model-free formalism. Conformational exchange observed in RBD1, which is absent in RBD2, may correlate to the mechanism of RNA binding.
- Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, Missouri 63110, USA.
Organizational Affiliation: