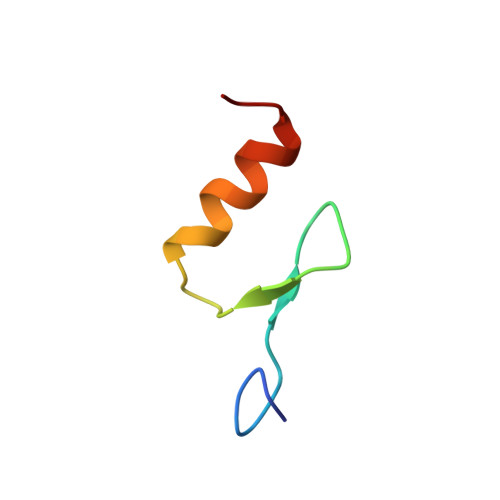
Solution structures of the DNA-binding domains of immune-related zinc-finger protein ZFAT
Tochio, N., Umehara, T., Nakabayashi, K., Yoneyama, M., Tsuda, K., Shirouzu, M., Koshiba, S., Watanabe, S., Kigawa, T., Sasazuki, T., Shirasawa, S., Yokoyama, S.(2015) J Struct Funct Genomics 16: 55-65
- PubMed: 25801860 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s10969-015-9196-3
- Primary Citation Related Structures:
2RUT, 2RUU, 2RUV, 2RUW, 2RUX, 2RUY, 2RUZ, 2RV0, 2RV1, 2RV2, 2RV3, 2RV4, 2RV5, 2RV6, 2RV7 - PubMed Abstract:
ZFAT is a transcriptional regulator, containing eighteen C2H2-type zinc-fingers and one AT-hook, involved in autoimmune thyroid disease, apoptosis, and immune-related cell survival. We determined the solution structures of the thirteen individual ZFAT zinc-fingers (ZF) and the tandemly arrayed zinc-fingers in the regions from ZF2 to ZF5, by NMR spectroscopy. ZFAT has eight uncommon bulged-out helix-containing zinc-fingers, and six of their structures (ZF4, ZF5, ZF6, ZF10, ZF11, and ZF13) were determined. The distribution patterns of the putative DNA-binding surface residues are different among the ZFAT zinc-fingers, suggesting the distinct DNA sequence preferences of the N-terminal and C-terminal zinc-fingers. Since ZFAT has three to five consecutive tandem zinc-fingers, which may cooperatively function as a unit, we also determined two tandemly arrayed zinc-finger structures, between ZF2 to ZF4 and ZF3 to ZF5. Our NMR spectroscopic analysis detected the interaction between ZF4 and ZF5, which are connected by an uncommon linker sequence, KKIK. The ZF4-ZF5 linker restrained the relative structural space between the two zinc-fingers in solution, unlike the other linker regions with determined structures, suggesting the involvement of the ZF4-ZF5 interfinger linker in the regulation of ZFAT function.
- RIKEN Systems and Structural Biology Center, 1-7-22 Suehiro-cho, Tsurumi, Yokohama, 230-0045 Japan.
Organizational Affiliation: