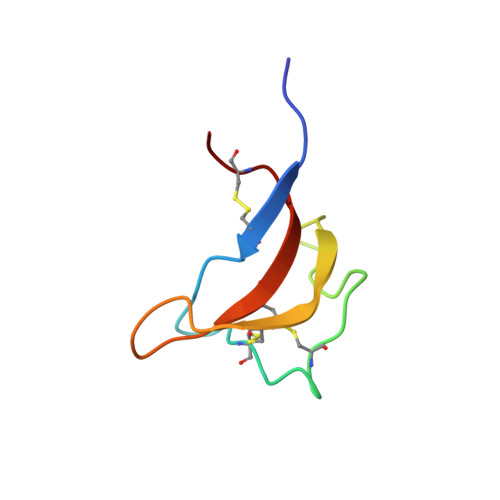
Solution structure of the rat P2X4 receptor head domain involved in inhibitory metal binding
Igawa, T., Abe, Y., Tsuda, M., Inoue, K., Ueda, T.(2015) FEBS Lett 589: 680-686
- PubMed: 25662851 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2015.01.034
- Primary Citation Related Structures:
2RUP - PubMed Abstract:
The P2X receptor is an ATP-gated cation channel expressed on the plasma membrane. The head domain (Gln111-Val167 in the rat P2X4 receptor) regulates ATP-induced cation influx. In this study, we prepared a head domain with three disulfide bonds, such as the intact rat P2X4 receptor contains. NMR analysis showed that the head domain possessed the same fold as in the zebrafish P2X4 receptor previously determined by crystallography. Furthermore, the inhibitory, divalent, metal ion binding sites were determined by NMR techniques. These findings will be useful for the design of specific inhibitors for the P2X receptor family.
- Department of Protein Structure, Function and Design, Graduate School of Pharmaceutical Sciences, Kyushu University, Fukuoka, Japan.
Organizational Affiliation: