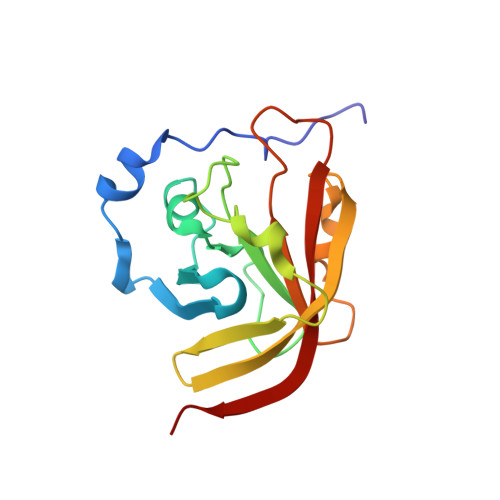
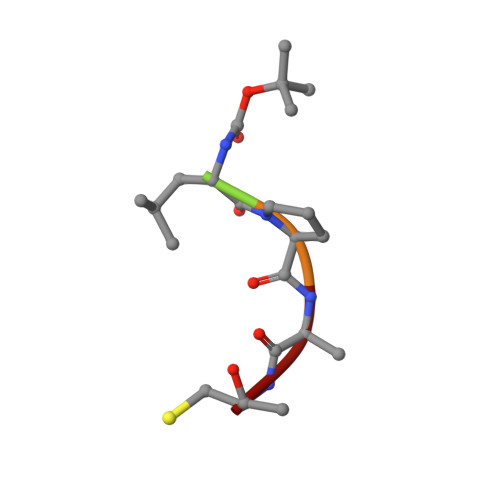
Structure of the Bacillus anthracis Sortase A Enzyme Bound to Its Sorting Signal: A FLEXIBLE AMINO-TERMINAL APPENDAGE MODULATES SUBSTRATE ACCESS.
Chan, A.H., Yi, S.W., Terwilliger, A.L., Maresso, A.W., Jung, M.E., Clubb, R.T.(2015) J Biological Chem 290: 25461-25474
- PubMed: 26324714 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.670984
- Primary Citation Related Structures:
2RUI - PubMed Abstract:
The endospore forming bacterium Bacillus anthracis causes lethal anthrax disease in humans and animals. The ability of this pathogen to replicate within macrophages is dependent upon the display of bacterial surface proteins attached to the cell wall by the B. anthracis Sortase A ((Ba)SrtA) enzyme. Previously, we discovered that the class A (Ba)SrtA sortase contains a unique N-terminal appendage that wraps around the body of the protein to contact the active site of the enzyme. To gain insight into its function, we determined the NMR structure of (Ba)SrtA bound to a LPXTG sorting signal analog. The structure, combined with dynamics, kinetics, and whole cell protein display data suggest that the N terminus modulates substrate access to the enzyme. We propose that it may increase the efficiency of protein display by reducing the unproductive hydrolytic cleavage of enzyme-protein covalent intermediates that form during the cell wall anchoring reaction. Notably, a key active site loop (β7/β8 loop) undergoes a disordered to ordered transition upon binding the sorting signal, potentially facilitating recognition of lipid II.
- From the Department of Chemistry and Biochemistry, UCLA-DOE Institute of Genomics and Proteomics, and the Molecular Biology Institute, University of California, Los Angeles, California 90095 and.
Organizational Affiliation: