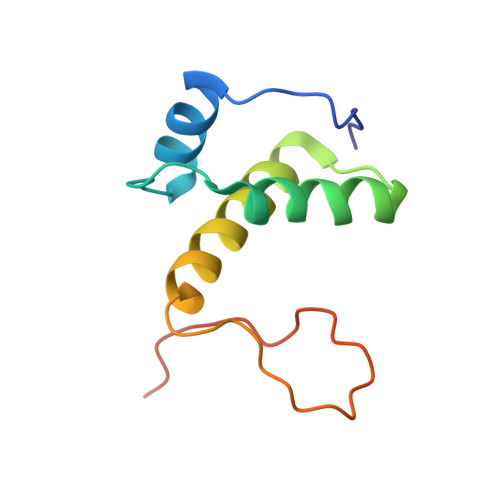
Structure and mechanism of the primosome protein DnaT-functional structures for homotrimerization, dissociation of ssDNA from the PriB·ssDNA complex, and formation of the DnaT·ssDNA complex.
Fujiyama, S., Abe, Y., Tani, J., Urabe, M., Sato, K., Aramaki, T., Katayama, T., Ueda, T.(2014) FEBS J 281: 5356-5370
- PubMed: 25265331 Search on PubMed
- DOI: https://doi.org/10.1111/febs.13080
- Primary Citation Related Structures:
2RU8 - PubMed Abstract:
In Escherichia coli, the primosome plays an essential role in replication restart after dissociation of replisomes at the damaged replication fork. As well as PriA and PriB, DnaT, an ssDNA-binding protein, is a key member of the primosome. In this study, limited proteolysis indicated that E. coli DnaT was composed of two domains, consistent with the results of recent studies using Klebsiella pneumonia DnaT. We also found that a specific 24-residue region (Phe42-Asp66) in the N-terminal domain (1-88) was crucial for DnaT trimerization. Moreover, we determined the structure of the DnaT C-terminal domain (89-179) by NMR spectroscopy. This domain included three α-helices and a long flexible C-terminal tail, similar to the C-terminal subdomain of the AAA+ ATPase family. The neighboring histidines, His136 and His137, at a position corresponding to the AAA+ sensor II motif, were suggested to form an ssDNA-binding site. Furthermore, we found that the acidic linker between the two domains had an activity for dissociating ssDNA from the PriB·ssDNA complexes in a manner supported by the conserved acidic residues Asp70 and Glu76. Thus, these findings provide a novel structural basis for understanding the mechanism of DnaT in exposure of ssDNA and reloading of the replicative helicase at the stalled replication fork. The coordinates used for the ensemble of NMR structures have been deposited in the Protein Data Bank under accession code 2ru8. The NMR data have been deposited in the BioMagResBank (www.bmrb.wisc.edu) under accession number 11549.
- Department of Protein Structure, Function and Design, Graduate School of Pharmaceutical Sciences, Kyushu University, Fukuoka, Japan.
Organizational Affiliation: