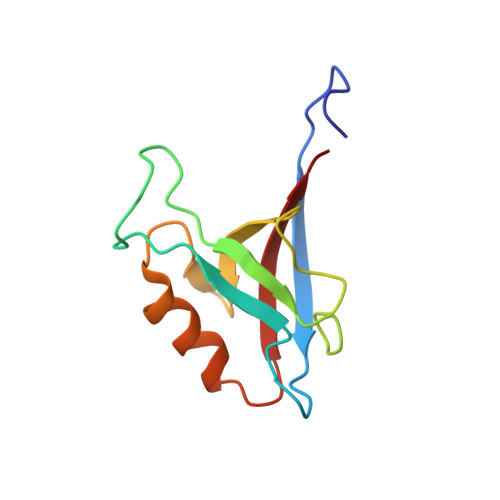
1H, 13C, and 15N resonance assignment of the first PDZ domain of mouse ZO-1
Umetsu, Y., Goda, N., Taniguchi, R., Satomura, K., Ikegami, T., Furuse, M., Hiroaki, H.(2011) Biomol NMR Assign 5: 207-210
- PubMed: 21431884 Search on PubMed
- DOI: https://doi.org/10.1007/s12104-011-9301-x
- Primary Citation Related Structures:
2RRM - PubMed Abstract:
Zonula occludens-1 (ZO-1) is a scaffolding molecule critical to the formation of intercellular adhesion structures, such as tight junctions (TJs) and adherens junctions (AJs). ZO-1 contains three PDZ domains followed by a GUK domain and a ZU5 domain. The first PDZ of ZO-1 (ZO-1(PDZ1)) serves as a protein-protein interaction module and interacts with the C-termini of almost all claudins to initiate the formation of a belt-like structure on the lateral membranes, thereby promoting TJ formation. It has been recently reported that approximately 15% of all PDZ domains bind phosphoinositides, and ZO-1(PDZ1) is the one of these. Here we report the (15)N, (13)C, and (1)H chemical shift assignments of the first PDZ domain of mouse ZO-1. The resonance assignments obtained in this work may contribute in clarifying the interplay between the two binary interactions, ZO-1(PDZ1)-claudins and ZO-1(PDZ1)-phospholipids, and suggesting a novel regulation mechanism underlying the formation and maintenance of cell-cell adhesion machinery downstream of the phospholipid signaling pathways.
- Division of Structural Biology Graduate School of Medicine, Kobe University, Kobe, Hyogo, Japan.
Organizational Affiliation: