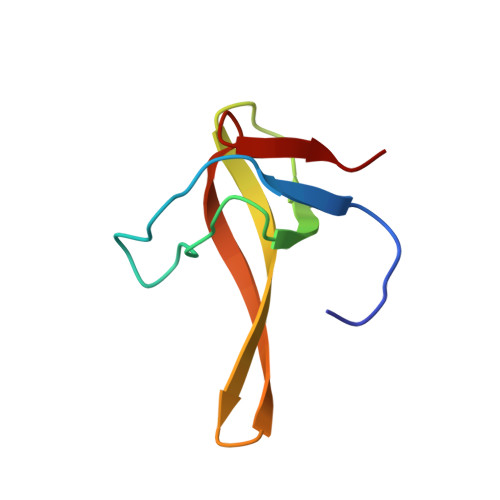
Solution structure and dynamics of the chimeric SH3 domains, SHH- and SHA-"Bergeracs".
Kutyshenko, V.P., Prokhorov, D.A., Timchenko, M.A., Kudrevatykh, Y.A., Gushchina, L.V., Khristoforov, V.S., Filimonov, V.V., Uversky, V.N.(2009) Biochim Biophys Acta 1794: 1813-1822
- PubMed: 19732853
- DOI: https://doi.org/10.1016/j.bbapap.2009.08.021
- Primary Citation Related Structures:
2ROT - PubMed Abstract:
Two chimeric proteins, SHcapital EN, Cyrillic and SHA of the "SH3-Bergerac" family (where the beta-turn N47D48 in spectrin SH3 domain was substituted for KITVNGKTYE or KATANGKTYE sequences, respectively), were analyzed by high-resolution NMR to resolve their spatial structures and to analyze their dynamics. Although the presence of a stable beta-hairpin in the region of the insertion was confirmed, the introduced extension of the polypeptide chain in SHcapital EN, Cyrillic (approximately 17%) practically did not affect the total molecule topology. Interestingly, the introduced beta-hairpin had higher mobility in comparison with other protein regions. Finally, we performed a disorder prediction with the PONDR VSL2 algorithm and discovered that the inserted beta-hairpin in both SHH and SHA proteins exhibited significant propensity for intrinsic disorder and therefore for high mobility. In agreement with the experimental data, the predisposition for the increased intramolecular mobility was noticeably higher in SHA.
- Institute of Theoretical and Experimental Biophysics of Russian Academy of Science, 142290 Pushchino, Moscow Region, Russia. kutyshenko@rambler.ru
Organizational Affiliation: