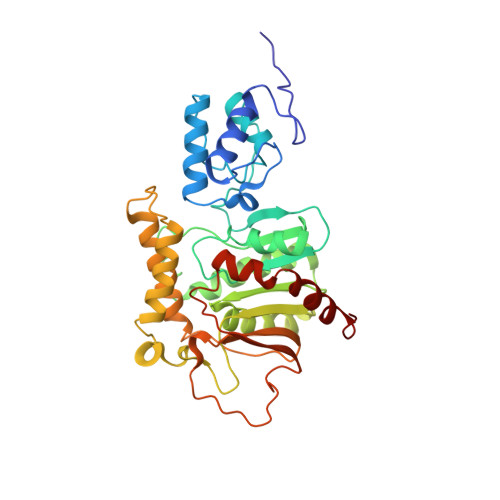
Dynamic thiolation-thioesterase structure of a non-ribosomal peptide synthetase
Frueh, D.P., Arthanari, H., Koglin, A., Vosburg, D.A., Bennett, A.E., Walsh, C.T., Wagner, G.(2008) Nature 454: 903-906
- PubMed: 18704088 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature07162
- Primary Citation Related Structures:
2ROQ - PubMed Abstract:
Non-ribosomal peptide synthetases (NRPS) and polyketide synthases (PKS) produce numerous secondary metabolites with various therapeutic/antibiotic properties. Like fatty acid synthases (FAS), these enzymes are organized in modular assembly lines in which each module, made of conserved domains, incorporates a given monomer unit into the growing chain. Knowledge about domain or module interactions may enable reengineering of this assembly line enzymatic organization and open avenues for the design of new bioactive compounds with improved therapeutic properties. So far, little structural information has been available on how the domains interact and communicate. This may be because of inherent interdomain mobility hindering crystallization, or because crystallized molecules may not represent the active domain orientations. In solution, the large size and internal dynamics of multidomain fragments (>35 kilodaltons) make structure determination by nuclear magnetic resonance a challenge and require advanced technologies. Here we present the solution structure of the apo-thiolation-thioesterase (T-TE) di-domain fragment of the Escherichia coli enterobactin synthetase EntF NRPS subunit. In the holoenzyme, the T domain carries the growing chain tethered to a 4'-phosphopantetheine whereas the TE domain catalyses hydrolysis and cyclization of the iron chelator enterobactin. The T-TE di-domain forms a compact but dynamic structure with a well-defined domain interface; the two active sites are at a suitable distance for substrate transfer from T to TE. We observe extensive interdomain and intradomain motions for well-defined regions and show that these are modulated by interactions with proteins that participate in the biosynthesis. The T-TE interaction described here provides a model for NRPS, PKS and FAS function in general as T-TE-like di-domains typically catalyse the last step in numerous assembly-line chain-termination machineries.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, Massachusetts 02115, USA. dominique_frueh@hms.harvard.edu
Organizational Affiliation: