Chemical synthesis and solution structure of a spider toxin that affects the inactivation of mammalian sodium channels
Yamaji, N., Nishio, H., Villegas, E., Nishiuchi, Y., Corzo, G.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Neurotoxin magi-4 | 43 | N/A | Mutation(s): 0 | 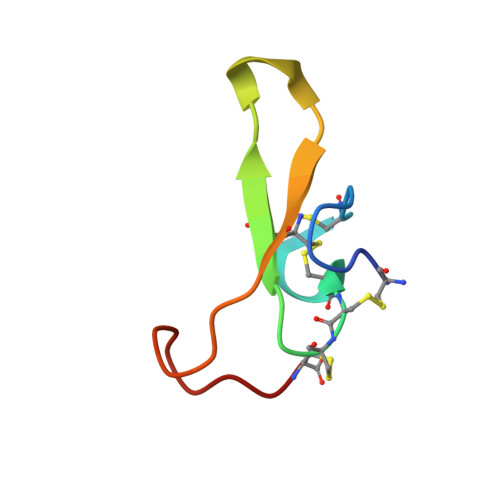 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P83560 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||