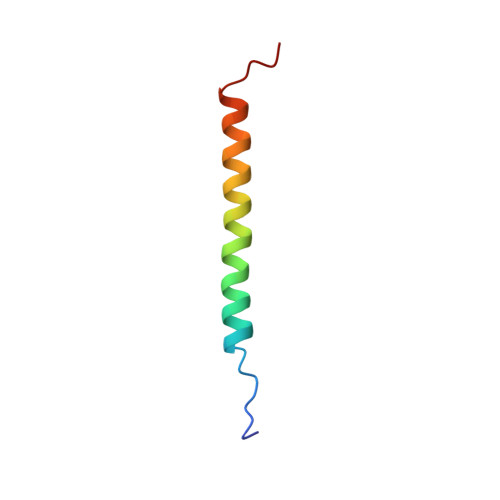
Structure of the Integrin beta3 Transmembrane Segment in Phospholipid Bicelles and Detergent Micelles
Lau, T.L., Partridge, A.W., Ginsberg, M.H., Ulmer, T.S.(2008) Biochemistry 47: 4008-4016
- PubMed: 18321071 Search on PubMed
- DOI: https://doi.org/10.1021/bi800107a
- Primary Citation Related Structures:
2RMZ, 2RN0 - PubMed Abstract:
Integrin adhesion receptors transduce bidirectional signals across the plasma membrane, with the integrin transmembrane domains acting as conduits in this process. Here, we report the first high-resolution structure of an integrin transmembrane domain. To assess the influence of the membrane model system, structure determinations of the beta3 integrin transmembrane segment and flanking sequences were carried out in both phospholipid bicelles and detergent micelles. In bicelles, a 30-residue linear alpha-helix, encompassing residues I693-H772, is adopted, of which I693-I721 appear embedded in the hydrophobic bicelle core. This relatively long transmembrane helix implies a pronounced helix tilt within a typical lipid bilayer, which facilitates the snorkeling of K716's charged side chain out of the lipid core while simultaneously immersing hydrophobic L717-I721 in the membrane. A shortening of bicelle lipid hydrocarbon tails does not lead to the transfer of L717-I721 into the aqueous phase, suggesting that the reported embedding represents the preferred beta3 state. The nature of the lipid headgroup affected only the intracellular part of the transmembrane helix, indicating that an asymmetric lipid distribution is not required for studying the beta3 transmembrane segment. In the micelle, residues L717-I721 are also embedded but deviate from linear alpha-helical conformation in contrast to I693-K716, which closely resemble the bicelle structure.
- Department of Biochemistry and Molecular Biology and Zilkha Neurogenetic Institute, Keck School of Medicine, University of Southern California, 1501 San Pablo Street, Los Angeles, California 90033, USA.
Organizational Affiliation: