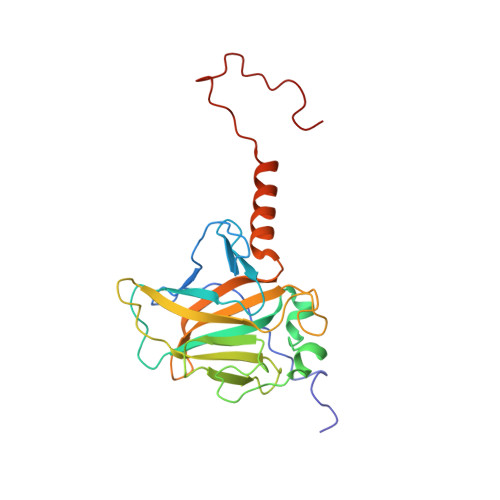
Solution structure and binding specificity of the p63 DNA binding domain
Enthart, A., Klein, C., Dehner, A., Coles, M., Gemmecker, G., Kessler, H., Hagn, F.(2016) Sci Rep 6: 26707-26707
- PubMed: 27225672 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep26707
- Primary Citation Related Structures:
2RMN - PubMed Abstract:
p63 is a close homologue of p53 and, together with p73, is grouped into the p53 family of transcription factors. p63 is known to be involved in the induction of controlled apoptosis important for differentiation processes, germ line integrity and development. Despite its high homology to p53, especially within the DNA binding domain (DBD), p63-DBD does not show cooperative DNA binding properties and is significantly more stable against thermal and chemical denaturation. Here, we determined the solution structure of p63-DBD and show that it is markedly less dynamic than p53-DBD. In addition, we also investigate the effect of a double salt bridge present in p53-DBD, but not in p63-DBD on the cooperative binding behavior and specificity to various DNA sites. Restoration of the salt bridges in p63-DBD by mutagenesis leads to enhanced binding affinity to p53-specific, but not p63-specific response elements. Furthermore, we show that p63-DBD is capable of binding to anti-apoptotic BclxL via its DNA binding interface, a feature that has only been shown for p53 so far. These data suggest that all p53 family members - despite alterations in the specificity and binding affinity - are capable of activating pro-apoptotic pathways in a tissue specific manner.
- Center for Integrated Protein Science Munich (CIPSM) at the Department Chemie, Technische Universität München, 85747 Garching, Germany.
Organizational Affiliation: