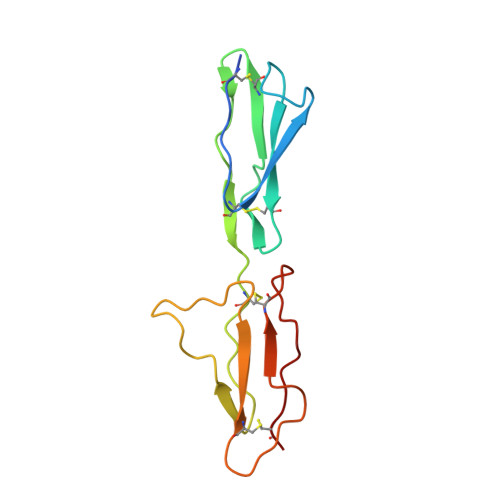
Structure of the N-terminal region of complement factor H and conformational implications of disease-linked sequence variations.
Hocking, H.G., Herbert, A.P., Kavanagh, D., Soares, D.C., Ferreira, V.P., Pangburn, M.K., Uhrin, D., Barlow, P.N.(2008) J Biological Chem 283: 9475-9487
- PubMed: 18252712
- DOI: https://doi.org/10.1074/jbc.M709587200
- Primary Citation of Related Structures:
2RLP, 2RLQ - PubMed Abstract:
Factor H is a regulatory glycoprotein of the complement system. We expressed the three N-terminal complement control protein modules of human factor H (FH1-3) and confirmed FH1-3 to be the minimal unit with cofactor activity for C3b proteolysis by factor I. We reconstructed FH1-3 from NMR-derived structures of FH1-2 and FH2-3 revealing an approximately 105-A-long rod-like arrangement of the modules. In structural comparisons with other C3b-engaging proteins, factor H module 3 most closely resembles factor B module 3, consistent with factor H competing with factor B for binding C3b. Factor H modules 1, 2, and 3 each has a similar backbone structure to first, second, and third modules, respectively, of functional sites in decay accelerating factor and complement receptor type 1; the equivalent intermodular tilt and twist angles are also broadly similar. Resemblance between molecular surfaces is closest for first modules but absent in the case of second modules. Substitution of buried Val-62 with Ile (a factor H single nucleotide polymorphism potentially protective for age-related macular degeneration and dense deposit disease) causes rearrangements within the module 1 core and increases thermal stability but does not disturb the interface with module 2. Replacement of partially exposed (in module 1) Arg-53 by His (an atypical hemolytic uremic syndrome-linked mutation) did not impair structural integrity at 37 degrees C, but this FH1-2 mutant was less stable at higher temperatures; furthermore, chemical shift differences indicated potential for small structural changes at the module 1-2 interface.
- Edinburgh Biomolecular NMR Unit, Schools of Chemistry and Biological Sciences, Joseph Black Chemistry Bldg., University of Edinburgh, West Mains Road, Edinburgh, United Kingdom.
Organizational Affiliation: