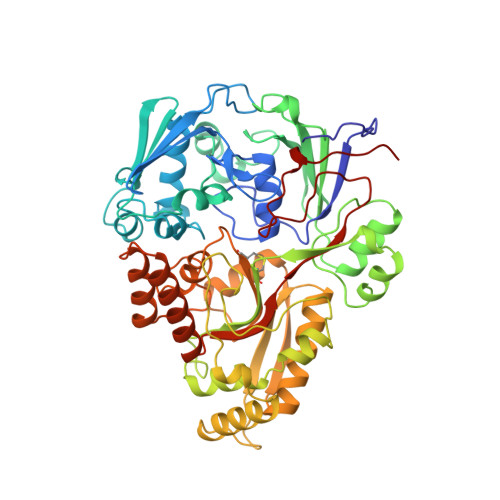
Peptide binding in OppA, the crystal structures of the periplasmic oligopeptide binding protein in the unliganded form and in complex with lysyllysine.
Sleigh, S.H., Tame, J.R., Dodson, E.J., Wilkinson, A.J.(1997) Biochemistry 36: 9747-9758
- PubMed: 9245406 Search on PubMed
- DOI: https://doi.org/10.1021/bi970457u
- Primary Citation Related Structures:
1RKM, 2RKM - PubMed Abstract:
The periplasmic oligopeptide binding protein, OppA, acts as the initial receptor for the uptake of peptides by the oligopeptide permease (Opp) in Gram-negative bacteria. Opp will handle peptides between two and five amino acid residues regardless of their sequence. The crystal structures of a series of OppA-peptide complexes have revealed an enclosed but versatile peptide binding pocket and have illustrated how tri- and tetrapeptide ligands are accommodated. Here, the crystal structures of (i) OppA complexed with a dipeptide (lysyllysine) and (ii) unliganded OppA have been solved using X-ray data extending to 1.8 and 2.4 A spacing, respectively. In the dipeptide complex, the alpha-amino group of the ligand is anchored through an ion pair interaction with Asp419, as observed in complexes with longer peptides. However, its alpha-carboxylate group forms water-mediated interactions with the guanidinium groups of Arg404 and Arg413 rather than the direct salt bridges to Arg413 and His371 observed in the tripeptide and tetrapeptide complexes, respectively. Isothermal titration calorimetric measurements of the binding of lysine-containing peptides of different lengths to OppA show that the dipeptide, KK, is bound with approximately 60-fold lower affinity than related tri- and tetrapeptides (KKK and KKKA, respectively). These data are discussed with reference to the calculated enthalpic and entropic contributions to ligand binding and the structures of the OppA peptide complexes. In the unliganded molecule, domain III has rotated as a rigid body through 26 degrees away from domains I and II, exposing the ligand binding site. The water structure in the binding cleft shows similarities to that in the various OppA-peptide complexes.
- Department of Chemistry, University of York, U.K.
Organizational Affiliation: