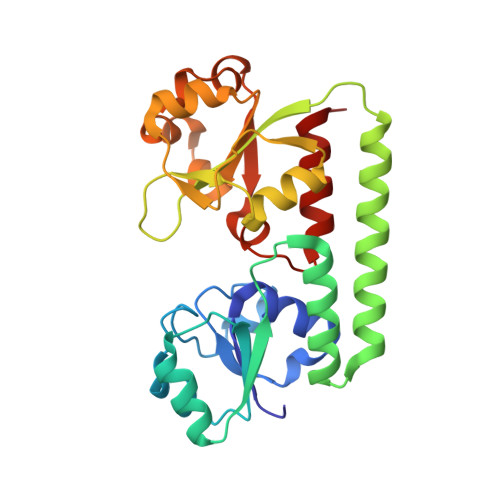
Holo- and apo-bound structures of bacterial periplasmic heme-binding proteins.
Ho, W.W., Li, H., Eakanunkul, S., Tong, Y., Wilks, A., Guo, M., Poulos, T.L.(2007) J Biological Chem 282: 35796-35802
- PubMed: 17925389 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M706761200
- Primary Citation Related Structures:
2R79, 2R7A, 2RG7 - PubMed Abstract:
An essential component of heme transport in Gram-negative bacterial pathogens is the periplasmic protein that shuttles heme between outer and inner membranes. We have solved the first crystal structures of two such proteins, ShuT from Shigella dysenteriae and PhuT from Pseudomonas aeruginosa. Both share a common architecture typical of Class III periplasmic binding proteins. The heme binds in a narrow cleft between the N- and C-terminal binding domains and is coordinated by a Tyr residue. A comparison of the heme-free (apo) and -bound (holo) structures indicates little change in structure other than minor alterations in the heme pocket and movement of the Tyr heme ligand from an "in" position where it can coordinate the heme iron to an "out" orientation where it points away from the heme pocket. The detailed architecture of the heme pocket is quite different in ShuT and PhuT. Although Arg(228) in PhuT H-bonds with a heme propionate, in ShuT a peptide loop partially takes up the space occupied by Arg(228), and there is no Lys or Arg H-bonding with the heme propionates. A comparison of PhuT/ShuT with the vitamin B(12)-binding protein BtuF and the hydroxamic-type siderophore-binding protein FhuD, the only two other structurally characterized Class III periplasmic binding proteins, demonstrates that PhuT/ShuT more closely resembles BtuF, which reflects the closer similarity in ligands, heme and B(12), compared with ligands for FhuD, a peptide siderophore.
- Department of Molecular Biology and Biochemistry, University of California, Irvine, CA 92697-3900, USA.
Organizational Affiliation: