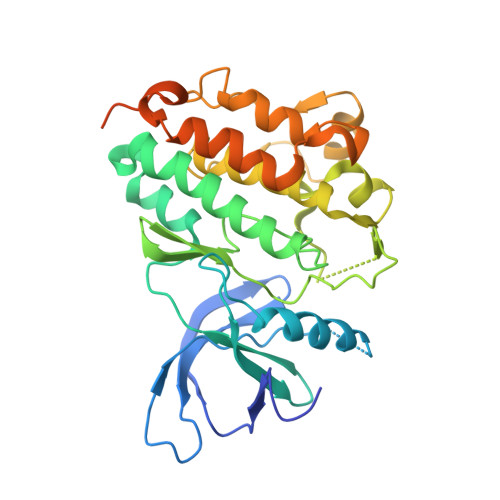
Inhibition of the EGF receptor by binding of MIG6 to an activating kinase domain interface.
Zhang, X., Pickin, K.A., Bose, R., Jura, N., Cole, P.A., Kuriyan, J.(2007) Nature 450: 741-744
- PubMed: 18046415 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature05998
- Primary Citation Related Structures:
2RF9, 2RFD, 2RFE - PubMed Abstract:
Members of the epidermal growth factor receptor family (EGFR/ERBB1, ERBB2/HER2, ERBB3/HER3 and ERBB4/HER4) are key targets for inhibition in cancer therapy. Critical for activation is the formation of an asymmetric dimer by the intracellular kinase domains, in which the carboxy-terminal lobe (C lobe) of one kinase domain induces an active conformation in the other. The cytoplasmic protein MIG6 (mitogen-induced gene 6; also known as ERRFI1) interacts with and inhibits the kinase domains of EGFR and ERBB2 (refs 3-5). Crystal structures of complexes between the EGFR kinase domain and a fragment of MIG6 show that a approximately 25-residue epitope (segment 1) from MIG6 binds to the distal surface of the C lobe of the kinase domain. Biochemical and cell-based analyses confirm that this interaction contributes to EGFR inhibition by blocking the formation of the activating dimer interface. A longer MIG6 peptide that is extended C terminal to segment 1 has increased potency as an inhibitor of the activated EGFR kinase domain, while retaining a critical dependence on segment 1. We show that signalling by EGFR molecules that contain constitutively active kinase domains still requires formation of the asymmetric dimer, underscoring the importance of dimer interface blockage in MIG6-mediated inhibition.
- Department of Molecular and Cell Biology, and Howard Hughes Medical Institute, California Institute for Quantitative Sciences, University of California, Berkeley, California 94720, USA.
Organizational Affiliation: