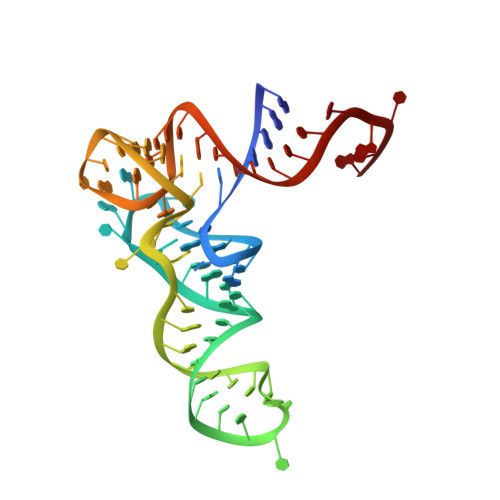
A rationally engineered misacylating aminoacyl-tRNA synthetase.
Bullock, T.L., Rodriguez-Hernandez, A., Corigliano, E.M., Perona, J.J.(2008) Proc Natl Acad Sci U S A 105: 7428-7433
- PubMed: 18477696 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0711812105
- Primary Citation Related Structures:
2RD2, 2RE8 - PubMed Abstract:
Information transfer from nucleic acid to protein is mediated by aminoacyl-tRNA synthetases, which catalyze the specific pairings of amino acids with transfer RNAs. Despite copious sequence and structural information on the 22 tRNA synthetase families, little is known of the enzyme signatures that specify amino acid selectivities. Here, we show that transplanting a conserved arginine residue from glutamyl-tRNA synthetase (GluRS) to glutaminyl-tRNA synthetase (GlnRS) improves the K(M) of GlnRS for noncognate glutamate. Two crystal structures of this C229R GlnRS mutant reveal that a conserved twin-arginine GluRS amino acid identity signature cannot be incorporated into GlnRS without disrupting surrounding protein structural elements that interact with the tRNA. Consistent with these findings, we show that cumulative replacement of other primary binding site residues in GlnRS, with those of GluRS, only slightly improves the ability of the GlnRS active site to accommodate glutamate. However, introduction of 22 amino acid replacements and one deletion, including substitution of the entire primary binding site and two surface loops adjacent to the region disrupted in C229R, improves the capacity of Escherichia coli GlnRS to synthesize misacylated Glu-tRNA(Gln) by 16,000-fold. This hybrid enzyme recapitulates the function of misacylating GluRS enzymes found in organisms that synthesize Gln-tRNA(Gln) by an alternative pathway. These findings implicate the RNA component of the contemporary GlnRS-tRNA(Gln) complex in mediating amino acid specificity. This role for tRNA may persist as a relic of primordial cells in which the evolution of the genetic code was driven by RNA-catalyzed amino acid-RNA pairing.
- Department of Chemistry and Biochemistry, and Interdepartmental Program in Biomolecular Science and Engineering, University of California, Santa Barbara, CA 93106-9510, USA.
Organizational Affiliation: