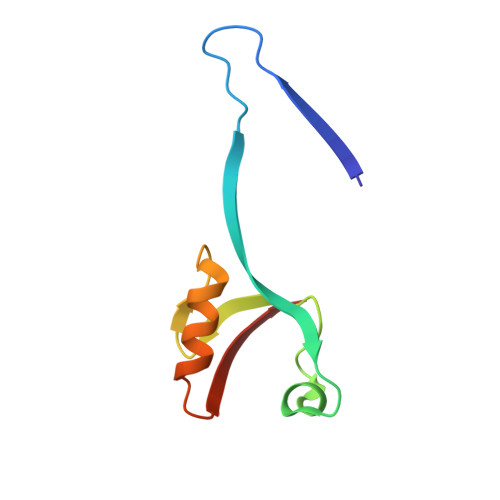
Domain swapping within PDZ2 is responsible for dimerization of ZO proteins.
Fanning, A.S., Lye, M.F., Anderson, J.M., Lavie, A.(2007) J Biological Chem 282: 37710-37716
- PubMed: 17928286 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M707255200
- Primary Citation Related Structures:
2RCZ - PubMed Abstract:
ZO-1 is a multidomain protein involved in cell-cell junctions and contains three PDZ domains, which are necessary for its function in vivo. PDZ domains play a central role in assembling diverse protein complexes through their ability to recognize short peptide motifs on other proteins. We determined the structure of the second of the three PDZ domains of ZO-1, which is known to promote dimerization as well as bind to C-terminal sequences on connexins. The dimer is stabilized by extensive symmetrical domain swapping of beta-strands, which is unlike any other known mechanism of PDZ dimerization. The canonical peptide-binding groove remains intact in both subunits of the PDZ2 dimer and is created by elements contributed from both monomers. This unique structure reveals an additional example of how PDZ domains dimerize and has multiple implications for both peptide binding and oligomerization in vivo.
- Department of Cell and Molecular Physiology, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599-7545, USA.
Organizational Affiliation: