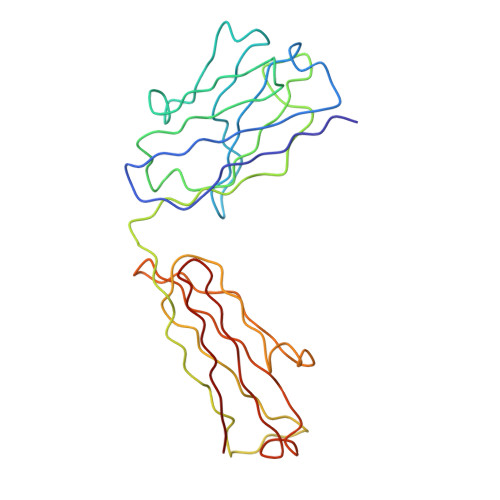
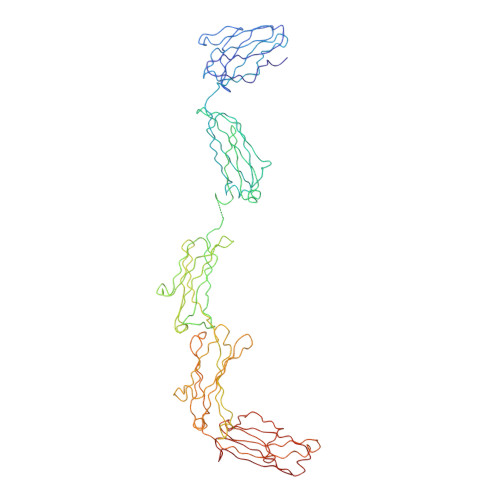
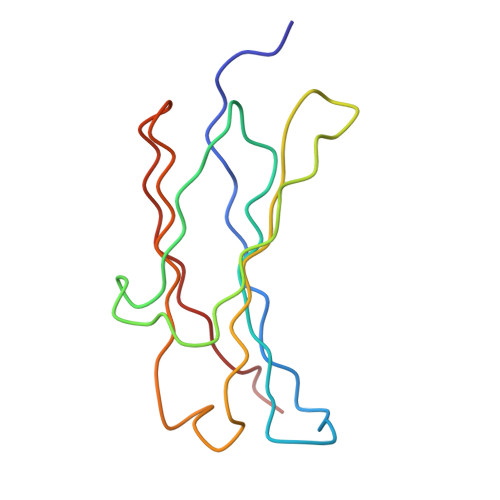
Solution structure of human and mouse immunoglobulin M by synchrotron X-ray scattering and molecular graphics modelling. A possible mechanism for complement activation.
Perkins, S.J., Nealis, A.S., Sutton, B.J., Feinstein, A.(1991) J Mol Biology 221: 1345-1366
- PubMed: 1942055 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(91)90937-2
- Primary Citation Related Structures:
2RCJ - PubMed Abstract:
The pentameric 71-domain structure of human and mouse immunoglobulin M (IgM) was investigated by synchrotron X-ray solution scattering and molecular graphics modelling. The radii of gyration RG of human IgM Quaife and its Fc5, IgM-S, Fab'2 and Fab fragments were determined as 12.2 nm, 6.1 nm, 6.1 nm, 4.9 nm and 2.9 nm in that order. The RG values were similar for mouse IgM P8 and its Fab'2 and Fab fragments, despite the presence of an additional carbohydrate site. The IgM scattering curves, to a nominal resolution of 5 nm, were compared with molecular graphics models based on published crystallographic alpha-carbon co-ordinates for the Fab and Fc structures of IgG. Good curve fits for Fab were obtained based on the crystal structure of Fab from IgG. A good curve fit was obtained for Fab'2, if the two Fab arms were positioned close together at their contact with the C mu 2 domains. The addition of the Fc fragment close to the C mu 2 domains of this Fab'2 model, to give a planar structure, accounted for the scattering curve of IgM-S. The Fc5 fragment was best modelled by a ring of five Fc monomers, constrained by packing considerations and disulphide bridge formation. A position for the J chain between two C mu 4 domains rather than at the centre of Fc5 was preferred. The intact IgM structure was best modelled using a planar arrangement of these Fab'2 and Fc5 models, with the side-to-side displacement of the Fab'2 arms in the plane of the IgM structure. All these models were consistent with hydrodynamic simulations of sedimentation data. The solution structure of IgM can therefore be reproduced quantitatively in terms of crystallographic structures for the fragments of IgG. Putative Clq binding sites have been identified on the C mu 3 domain. These would become accessible for interaction with Clq when the Fab'2 arms move out of the plane of the Fc5 disc in IgM, that is, a steric mechanism exposing pre-existing Clq sites. Comparison with a solution structure for Clq by neutron scattering shows that two or more of the six globular Clq heads in the hexameric head-and-stalk structure are readily able to make contacts with the putative Clq sites in the C mu 3 domains of free IgM if if the Clq arm-axis angle in solution is reduced from 40 degrees-45 degrees to 28 degrees. This could be the trigger for Cl activation.
- Department of Biochemistry and Chemistry, Royal Free Hospital School of Medicine, London, U.K.
Organizational Affiliation: