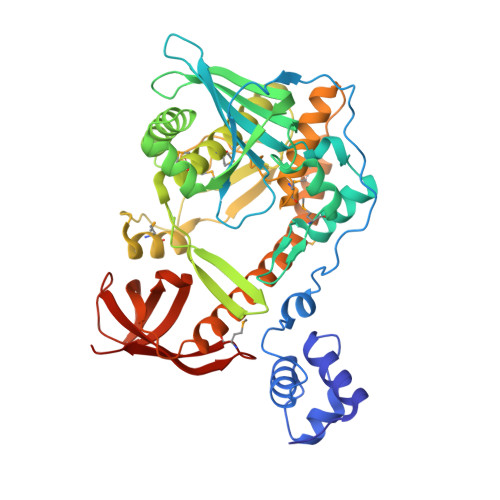
The Structure and Enzymatic Properties of a Novel RNase II Family Enzyme from Deinococcus radiodurans.
Schmier, B.J., Seetharaman, J., Deutscher, M.P., Hunt, J.F., Malhotra, A.(2012) J Mol Biology 415: 547-559
- PubMed: 22133431 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.11.031
- Primary Citation Related Structures:
2R7D, 2R7F - PubMed Abstract:
Exoribonucleases are vital in nearly all aspects of RNA metabolism, including RNA maturation, end-turnover, and degradation. RNase II and RNase R are paralogous members of the RNR superfamily of nonspecific, 3'→5', processive exoribonucleases. In Escherichia coli, RNase II plays a primary role in mRNA decay and has a preference for unstructured RNA. RNase R, in contrast, is capable of digesting structured RNA and plays a role in the degradation of both mRNA and stable RNA. Deinococcus radiodurans, a radiation-resistant bacterium, contains two RNR family members. The shorter of these, DrR63, includes a sequence signature typical of RNase R, but we show here that this enzyme is an RNase II-type exonuclease and cannot degrade structured RNA. We also report the crystal structure of this protein, now termed DrII. The DrII structure reveals a truncated RNA binding region in which the N-terminal cold shock domains, typical of most RNR family nucleases, are replaced by an unusual winged helix-turn-helix domain, where the "wing" is contributed by the C-terminal S1 domain. Consistent with its truncated RNA binding region, DrII is able to remove 3' overhangs from RNA molecules closer to duplexes than do other RNase II-type enzymes. DrII also displays distinct sensitivity to pyrimidine-rich regions of single-stranded RNA and is able to process tRNA precursors with adenosine-rich 3' extensions in vitro. These data indicate that DrII is the RNase II of D. radiodurans and that its structure and catalytic properties are distinct from those of other related enzymes.
- Department of Biochemistry and Molecular Biology, University of Miami Miller School of Medicine, PO Box 016129, Miami, FL 33101-6129, USA.
Organizational Affiliation: