Discovery of Dibenzo[c,f][2,7]naphthyridines as Potent and Selective 3-Phosphoinositide-Dependent Kinase-1 Inhibitors.
Gopalsamy, A., Shi, M., Boschelli, D.H., Williamson, R., Olland, A.M., Hu, Y., Krishnamurthy, G., Han, X., Arndt, K., Guo, B.(2007) J Med Chem 50: 5547-5549
- PubMed: 17941624
- DOI: https://doi.org/10.1021/jm070851i
- Primary Citation of Related Structures:
2R7B - PubMed Abstract:
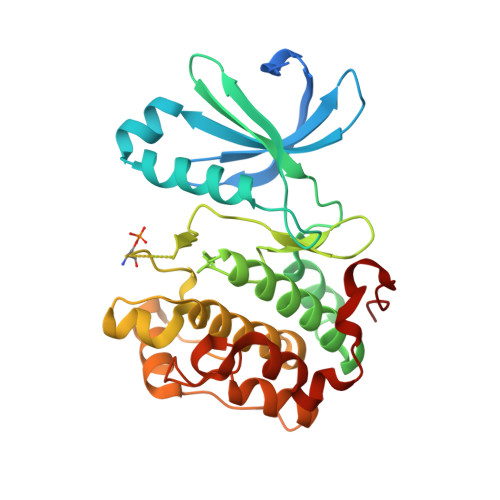
With high-throughput screening, substituted dibenzo[c,f][2,7]naphthyridine 1 was identified as a novel potent and selective phosphoinositide-dependent kinase-1 (PDK-1) inhibitor. Various regions of the lead molecule were explored to understand the SAR requirement for this scaffold. The crystal structure of 1 with kinase domain of PDK-1 confirmed the binding in the active site. The key interaction of the molecule with the active site residues, observed SAR, and the biological profile are discussed in detail.
- Chemical and Screening Sciences and Oncology, Wyeth Research, 401 N. Middletown Road, Pearl River, NY 10965, USA. gopalsa@wyeth.com
Organizational Affiliation: