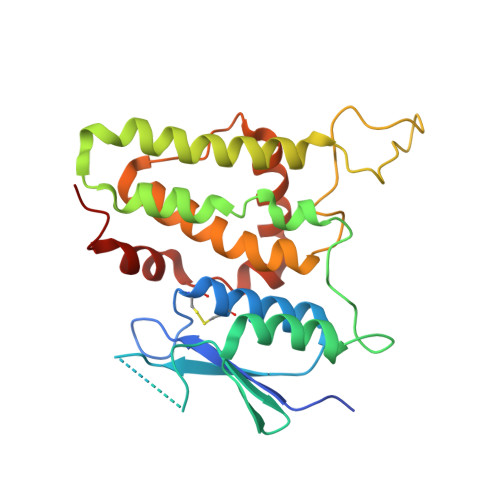
Structure of the Janus Protein Human CLIC2
Cromer, B.A., Gorman, M.A., Hansen, G., Adams, J.J., Coggan, M., Littler, D.R., Brown, L.J., Mazzanti, M., Breit, S.N., Curmi, P.M.G., Dulhunty, A.F., Board, P.G., Parker, M.W.(2007) J Mol Biology 374: 719-731
- PubMed: 17945253 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.09.041
- Primary Citation Related Structures:
2R4V, 2R5G - PubMed Abstract:
Chloride intracellular channel (CLIC) proteins possess the remarkable property of being able to convert from a water-soluble state to a membrane channel state. We determined the three-dimensional structure of human CLIC2 in its water-soluble form by X-ray crystallography at 1.8-A resolution from two crystal forms. In contrast to the previously characterized CLIC1 protein, which forms a possibly functionally important disulfide-induced dimer under oxidizing conditions, we show that CLIC2 possesses an intramolecular disulfide and that the protein remains monomeric irrespective of redox conditions. Site-directed mutagenesis studies show that removal of the intramolecular disulfide or introduction of cysteine residues in CLIC2, equivalent to those that form the intramolecular disulfide in CLIC1, does not cause dimer formation under oxidizing conditions. We also show that CLIC2 forms pH-dependent chloride channels in vitro with higher channel activity at low pH levels and that the channels are subject to redox regulation. In both crystal forms, we observed an extended loop region from the C-terminal domain, called the foot loop, inserting itself into an interdomain crevice of a neighboring molecule. The equivalent region in the structurally related glutathione transferase superfamily corresponds to the active site. This so-called foot-in-mouth interaction suggests that CLIC2 might recognize other proteins such as the ryanodine receptor through a similar interaction.
- Biota Structural Biology Laboratory, St. Vincent's Institute, Fitzroy, Victoria 3065, Australia.
Organizational Affiliation: