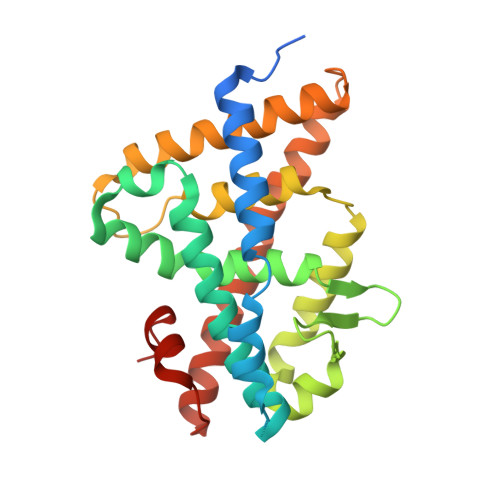
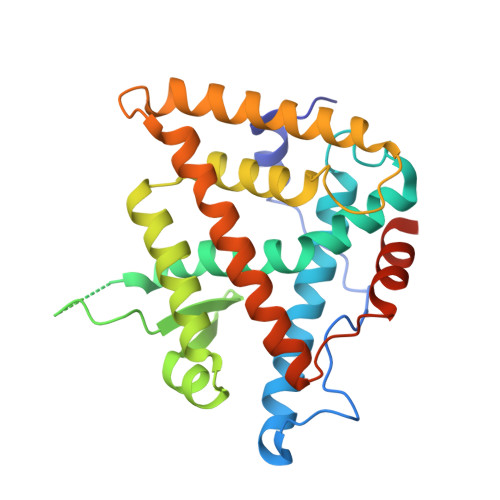
Critical Role of Desolvation in the Binding of 20-Hydroxyecdysone to the Ecdysone Receptor
Browning, C., Martin, E., Loch, C., Wurtz, J.M., Moras, D., Stote, R.H., Dejaegere, A.P., Billas, I.M.L.(2007) J Biological Chem 282: 32924-32934
- PubMed: 17848566 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M705559200
- Primary Citation Related Structures:
2R40 - PubMed Abstract:
The insect steroid hormone 20-hydroxyecdysone (20E) binds to its cognate nuclear receptor composed of the ecdysone receptor (EcR) and Ultraspiracle (USP) and triggers the main developmental transitions, in particular molting and metamorphosis. We present the crystal structure of the ligand-binding domains of EcR/USP in complex with 20E at 2.4A resolution and compare it with published structures of EcR/USP bound to ponasterone A (ponA). ponA is essentially identical to 20E but lacks the 25-OH group of 20E. The structure of 20E-bound EcR indicates that an additional hydrogen bond is formed compared with the ponA-bound receptor, yet, paradoxically, ponA has a significantly higher affinity for EcR than 20E. Theoretical studies based on docking and free energy methods lead to a rationale for understanding the difference in binding affinities between 20E and ponA. Results of the calculations indicate that the favorable contribution from the extra H-bond made by 25-OH of 20E is counterbalanced by its larger desolvation cost compared with that of ponA. The contribution of 25-OH to the binding affinity is further compared with those of 20- and 22-OH groups. Ligands that lack the 20- or 22-OH group are indeed known to bind less favorably to EcR than 20E, an effect opposite to that observed for ponA. The results indicate that their respective contributions to receptor-ligand complex stability reside mostly in their different contributions to solvation/desolvation. Together, the data demonstrate the critical role of ligand desolvation in determining binding affinity, with general implications for the binding of hormones to their cognate nuclear receptors.
- Département de Biologie et de Génomique Structurales, Institut de Génétique et de Biologie Moléculaire et Cellulaire, Ecole Supérieure de Biotechnologie de Strasbourg, Boulevard Sébastien Brant, Illkirch, France.
Organizational Affiliation: