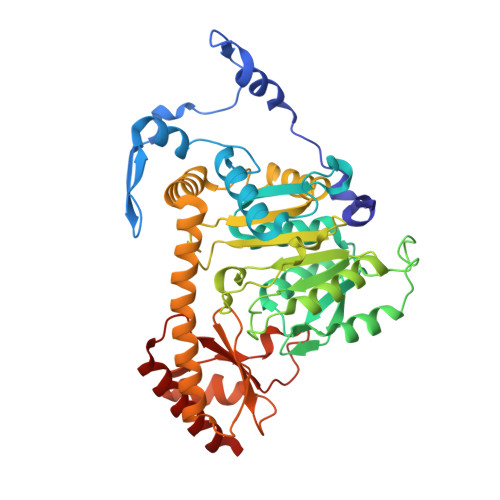
Crystal structure of human kynurenine aminotransferase II.
Han, Q., Robinson, H., Li, J.(2008) J Biological Chem 283: 3567-3573
- PubMed: 18056995 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M708358200
- Primary Citation Related Structures:
2QLR, 2R2N - PubMed Abstract:
Human kynurenine aminotransferase II (hKAT-II) efficiently catalyzes the transamination of knunrenine to kynurenic acid (KYNA). KYNA is the only known endogenous antagonist of N-methyl-D-aspartate (NMDA) receptors and is also an antagonist of 7-nicotinic acetylcholine receptors. Abnormal concentrations of brain KYNA have been implicated in the pathogenesis and development of several neurological and psychiatric diseases in humans. Consequently, enzymes involved in the production of brain KYNA have been considered potential regulatory targets. In this article, we report a 2.16 A crystal structure of hKAT-II and a 1.95 A structure of its complex with kynurenine. The protein architecture of hKAT-II reveals that it belongs to the fold-type I pyridoxal 5-phosphate (PLP)-dependent enzymes. In comparison with all subclasses of fold-type I-PLP-dependent enzymes, we propose that hKAT-II represents a novel subclass in the fold-type I enzymes because of the unique folding of its first 65 N-terminal residues. This study provides a molecular basis for future effort in maintaining physiological concentrations of KYNA through molecular and biochemical regulation of hKAT-II.
- Department of Biochemistry, Virginia Tech University, Blacksburg, Virginia 24061.
Organizational Affiliation: