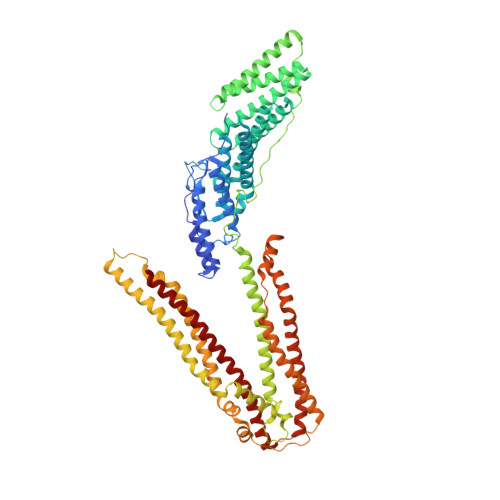
Structural and functional studies of ALIX interactions with YPX(n)L late domains of HIV-1 and EIAV.
Zhai, Q., Fisher, R.D., Chung, H.Y., Myszka, D.G., Sundquist, W.I., Hill, C.P.(2008) Nat Struct Mol Biol 15: 43-49
- PubMed: 18066081 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb1319
- Primary Citation Related Structures:
2R02, 2R03, 2R05 - PubMed Abstract:
Retrovirus budding requires short peptide motifs (late domains) located within the viral Gag protein that function by recruiting cellular factors. The YPX(n)L late domains of HIV and other lentiviruses recruit the protein ALIX (also known as AIP1), which also functions in vesicle formation at the multivesicular body and in the abscission stage of cytokinesis. Here, we report the crystal structures of ALIX in complex with the YPX(n)L late domains from HIV-1 and EIAV. The two distinct late domains bind at the same site on the ALIX V domain but adopt different conformations that allow them to make equivalent contacts. Binding studies and functional assays verified the importance of key interface residues and revealed that binding affinities are tuned by context-dependent effects. These results reveal how YPX(n)L late domains recruit ALIX to facilitate virus budding and how ALIX can bind YPX(n)L sequences with both n = 1 and n = 3.
- Department of Biochemistry, University of Utah School of Medicine, Salt Lake City, Utah 84112-5650, USA.
Organizational Affiliation: