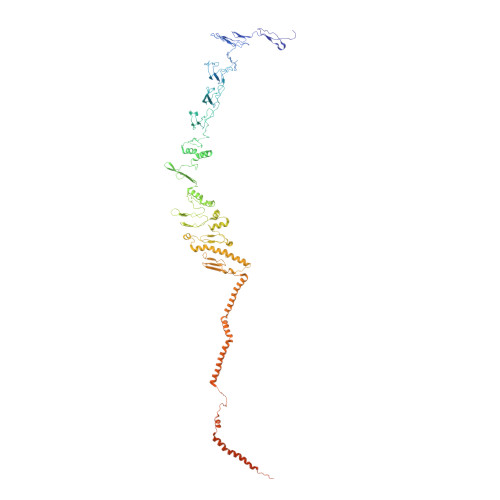
Draft crystal structure of the vault shell at 9-A resolution.
Anderson, D.H., Kickhoefer, V.A., Sievers, S.A., Rome, L.H., Eisenberg, D.(2007) PLoS Biol 5: e318-e318
- PubMed: 18044992 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.0050318
- Primary Citation Related Structures:
2QZV - PubMed Abstract:
Vaults are the largest known cytoplasmic ribonucleoprotein structures and may function in innate immunity. The vault shell self-assembles from 96 copies of major vault protein and encapsulates two other proteins and a small RNA. We crystallized rat liver vaults and several recombinant vaults, all among the largest non-icosahedral particles to have been crystallized. The best crystals thus far were formed from empty vaults built from a cysteine-tag construct of major vault protein (termed cpMVP vaults), diffracting to about 9-A resolution. The asymmetric unit contains a half vault of molecular mass 4.65 MDa. X-ray phasing was initiated by molecular replacement, using density from cryo-electron microscopy (cryo-EM). Phases were improved by density modification, including concentric 24- and 48-fold rotational symmetry averaging. From this, the continuous cryo-EM electron density separated into domain-like blocks. A draft atomic model of cpMVP was fit to this improved density from 15 domain models. Three domains were adapted from a nuclear magnetic resonance substructure. Nine domain models originated in ab initio tertiary structure prediction. Three C-terminal domains were built by fitting poly-alanine to the electron density. Locations of loops in this model provide sites to test vault functions and to exploit vaults as nanocapsules.
- Howard Hughes Medical Institute, University of California Los Angeles, Los Angeles, California, United States of America.
Organizational Affiliation: