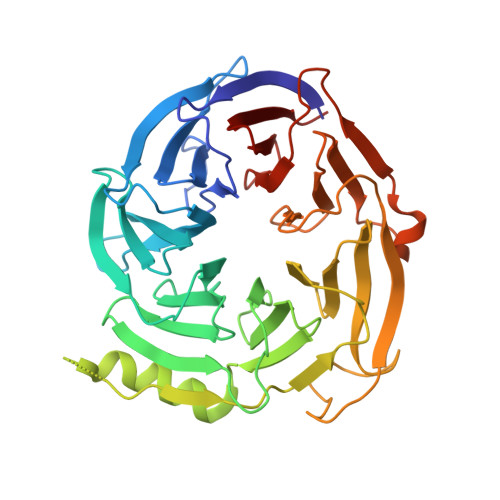
Structural basis of EZH2 recognition by EED
Han, Z., Xing, X., Hu, M., Zhang, Y., Liu, P., Chai, J.(2007) Structure 15: 1306-1315
- PubMed: 17937919 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2007.08.007
- Primary Citation Related Structures:
2QXV - PubMed Abstract:
The WD-repeat domain is a highly conserved recognition module in eukaryotes involved in diverse cellular processes. It is still not well understood how the bottom of a WD-repeat domain recognizes its binding partners. The WD-repeat-containing protein EED is one component of the PRC2 complex that possesses histone methyltransferase activity required for gene repression. Here we report the crystal structure of EED in complex with a 30 residue peptide from EZH2. The structure reveals that the peptide binds to the bottom of the WD-repeat domain of EED. The structural determinants of EZH2-EED interaction are present not only in EZH2 and EZH1 but also in its Drosophila homolog E(Z), suggesting that the recognition of ESC by E(Z) in Drosophila employs similar structural motifs. Structure-based mutagenesis identified critical residues from both EED and EZH2 for their interaction. The structure presented here may provide a template for understanding of how WD-repeat proteins recognize their interacting proteins.
- National Institute of Biological Sciences, Beijing 102206, China.
Organizational Affiliation: