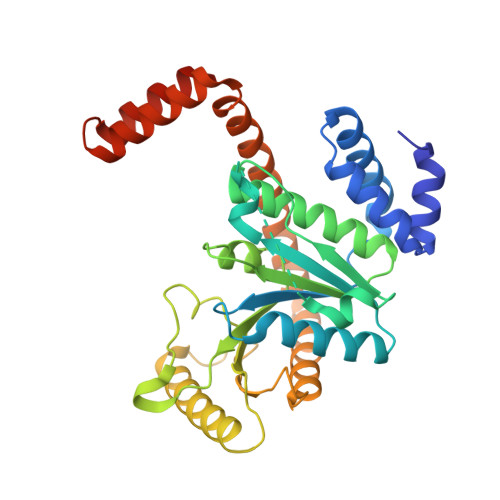
Crystal structure and mutagenesis of the metallochaperone MeaB: insight into the causes of methylmalonic aciduria.
Hubbard, P.A., Padovani, D., Labunska, T., Mahlstedt, S.A., Banerjee, R., Drennan, C.L.(2007) J Biological Chem 282: 31308-31316
- PubMed: 17728257 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M704850200
- Primary Citation Related Structures:
2QM7, 2QM8 - PubMed Abstract:
MeaB is an auxiliary protein that plays a crucial role in the protection and assembly of the B(12)-dependent enzyme methylmalonyl-CoA mutase. Impairments in the human homologue of MeaB, MMAA, lead to methylmalonic aciduria, an inborn error of metabolism. To explore the role of this metallochaperone, its structure was solved in the nucleotide-free form, as well as in the presence of product, GDP. MeaB is a homodimer, with each subunit containing a central alpha/beta-core G domain that is typical of the GTPase family, as well as alpha-helical extensions at the N and C termini that are not found in other metalloenzyme chaperone GTPases. The C-terminal extension appears to be essential for nucleotide-independent dimerization, and the N-terminal region is implicated in protein-protein interaction with its partner protein, methylmalonyl-CoA mutase. The structure of MeaB confirms that it is a member of the G3E family of P-loop GTPases, which contains other putative metallochaperones HypB, CooC, and UreG. Interestingly, the so-called switch regions, responsible for signal transduction following GTP hydrolysis, are found at the dimer interface of MeaB instead of being positioned at the surface of the protein where its partner protein methylmalonyl-CoA mutase should bind. This observation suggests a large conformation change of MeaB must occur between the GDP- and GTP-bound forms of this protein. Because of their high sequence homology, the missense mutations in MMAA that result in methylmalonic aciduria have been mapped onto MeaB and, in conjunction with mutagenesis data, provide possible explanations for the pathology of this disease.
- Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA.
Organizational Affiliation: