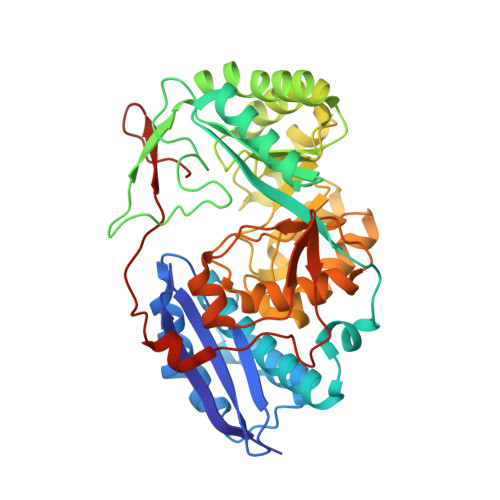
Evolution of enzymatic activities in the enolase superfamily: D-Mannonate dehydratase from Novosphingobium aromaticivorans.
Rakus, J.F., Fedorov, A.A., Fedorov, E.V., Glasner, M.E., Vick, J.E., Babbitt, P.C., Almo, S.C., Gerlt, J.A.(2007) Biochemistry 46: 12896-12908
- PubMed: 17944491
- DOI: https://doi.org/10.1021/bi701703w
- Primary Citation Related Structures:
2QJJ, 2QJM, 2QJN - PubMed Abstract:
The d-mannonate dehydratase (ManD) function was assigned to a group of orthologous proteins in the mechanistically diverse enolase superfamily by screening a library of acid sugars. Structures of the wild type ManD from Novosphingobium aromaticivorans were determined at pH 7.5 in the presence of Mg2+ and also in the presence of Mg2+ and the 2-keto-3-keto-d-gluconate dehydration product; the structure of the catalytically active K271E mutant was determined at pH 5.5 in the presence of the d-mannonate substrate. As previously observed in the structures of other members of the enolase superfamily, ManD contains two domains, an N-terminal alpha+beta capping domain and a (beta/alpha)7beta-barrel domain. The barrel domain contains the ligands for the essential Mg2+, Asp 210, Glu 236, and Glu 262, at the ends of the third, fourth, and fifth beta-strands of the barrel domain, respectively. However, the barrel domain lacks both the Lys acid/base catalyst at the end of the second beta-strand and the His-Asp dyad acid/base catalyst at the ends of the seventh and sixth beta-strands, respectively, that are found in many members of the superfamily. Instead, a hydrogen-bonded dyad of Tyr 159 in a loop following the second beta-strand and Arg 147 at the end of the second beta-strand are positioned to initiate the reaction by abstraction of the 2-proton. Both Tyr 159 and His 212, at the end of the third beta-strand, are positioned to facilitate both syn-dehydration and ketonization of the resulting enol intermediate to yield the 2-keto-3-keto-d-gluconate product with the observed retention of configuration. The identities and locations of these acid/base catalysts as well as of cationic amino acid residues that stabilize the enolate anion intermediate define a new structural strategy for catalysis (subgroup) in the mechanistically diverse enolase superfamily. With these differences, we provide additional evidence that the ligands for the essential Mg2+ are the only conserved residues in the enolase superfamily, establishing the primary functional importance of the Mg2+-assisted strategy for stabilizing the enolate anion intermediate.
- Department of Biochemistry, University of Illinois at Urbana-Champaign, 600 S. Mathews Avenue, Urbana, Illinois 61801, USA.
Organizational Affiliation: