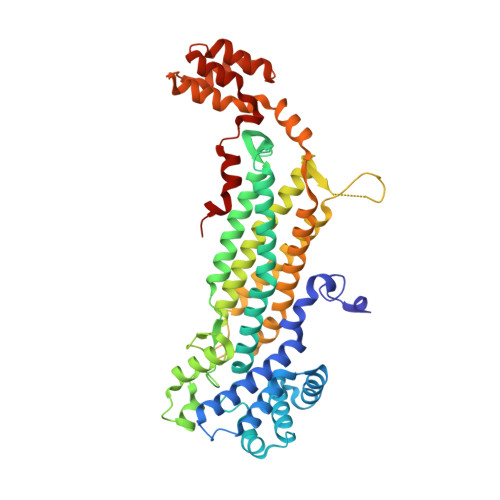
Plasmodium vivax adenylosuccinate lyase Pv003765 with AMP bound
Lunin, V.V., Wernimont, A.K., Lew, J., Kozieradzki, I., Bochkarev, A., Arrowsmith, C.H., Sundstrom, M., Weigelt, J., Edwards, A.E., Hui, R., Hills, T., Altamentova, S., Structural Genomics Consortium (SGC)To be published.