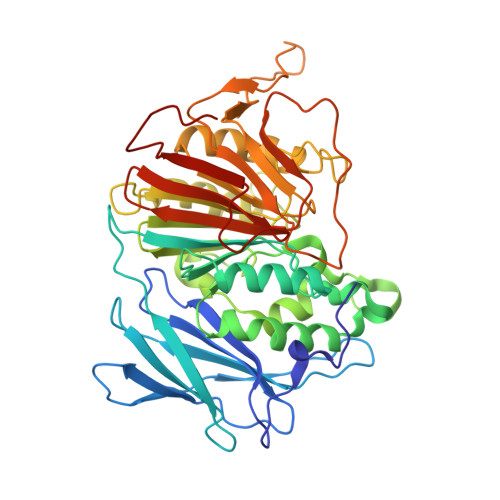
Crystal structures of a purple acid phosphatase, representing different steps of this enzyme's catalytic cycle.
Schenk, G., Elliott, T.W., Leung, E., Carrington, L.E., Mitic, N., Gahan, L.R., Guddat, L.W.(2008) BMC Struct Biol 8: 6-6
- PubMed: 18234116 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-8-6
- Primary Citation Related Structures:
2QFP, 2QFR - PubMed Abstract:
Purple acid phosphatases belong to the family of binuclear metallohydrolases and are involved in a multitude of biological functions, ranging from bacterial killing and bone metabolism in animals to phosphate uptake in plants. Due to its role in bone resorption purple acid phosphatase has evolved into a promising target for the development of anti-osteoporotic chemotherapeutics. The design of specific and potent inhibitors for this enzyme is aided by detailed knowledge of its reaction mechanism. However, despite considerable effort in the last 10 years various aspects of the basic molecular mechanism of action are still not fully understood. Red kidney bean purple acid phosphatase is a heterovalent enzyme with an Fe(III)Zn(II) center in the active site. Two new structures with bound sulfate (2.4 A) and fluoride (2.2 A) provide insight into the pre-catalytic phase of its reaction cycle and phosphorolysis. The sulfate-bound structure illustrates the significance of an extensive hydrogen bonding network in the second coordination sphere in initial substrate binding and orientation prior to hydrolysis. Importantly, both metal ions are five-coordinate in this structure, with only one nucleophilic mu-hydroxide present in the metal-bridging position. The fluoride-bound structure provides visual support for an activation mechanism for this mu-hydroxide whereby substrate binding induces a shift of this bridging ligand towards the divalent metal ion, thus increasing its nucleophilicity. In combination with kinetic, crystallographic and spectroscopic data these structures of red kidney bean purple acid phosphatase facilitate the proposal of a comprehensive eight-step model for the catalytic mechanism of purple acid phosphatases in general.
- School of Molecular and Microbial Sciences, The University of Queensland, St, Lucia, QLD 4072, Australia. schenk@uq.edu.au
Organizational Affiliation: