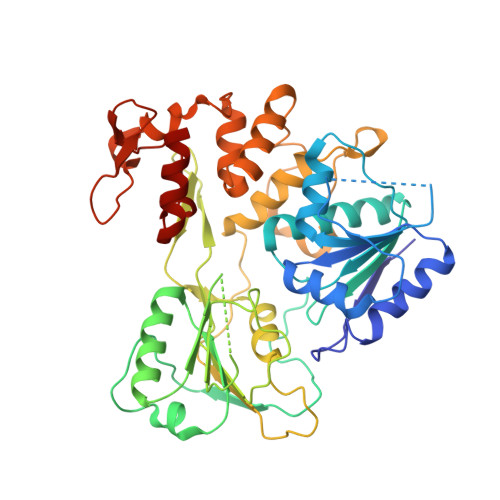
Crystal structure and activity of Kunjin virus NS3 helicase; protease and helicase domain assembly in the full length NS3 protein
Mastrangelo, E., Milani, M., Bollati, M., Selisko, B., Peyrane, F., Pandini, V., Sorrentino, G., Canard, B., Konarev, P.V., Svergun, D.I., de Lamballerie, X., Coutard, B., Khromykh, A.A., Bolognesi, M.(2007) J Mol Biology 372: 444-455
- PubMed: 17658551 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.06.055
- Primary Citation Related Structures:
2QEQ - PubMed Abstract:
Flaviviral NS3 is a multifunctional protein displaying N-terminal protease activity in addition to C-terminal helicase, nucleoside 5'-triphosphatase (NTPase), and 5'-terminal RNA triphosphatase (RTPase) activities. NS3 is held to support the separation of RNA daughter and template strands during viral replication. In addition, NS3 assists the initiation of replication by unwinding the RNA secondary structure in the 3' non-translated region (NTR). We report here the three-dimensional structure (at 3.1 A resolution) of the NS3 helicase domain (residues 186-619; NS3:186-619) from Kunjin virus, an Australian variant of the West Nile virus. As for homologous helicases, NS3:186-619 is composed of three domains, two of which are structurally related and held to host the NTPase and RTPase active sites. The third domain (C-terminal) is involved in RNA binding/recognition. The NS3:186-619 construct occurs as a dimer in solution and in the crystals. We show that NS3:186-619 displays both ATPase and RTPase activities, that it can unwind a double-stranded RNA substrate, being however inactive on a double-stranded DNA substrate. Analysis of different constructs shows that full length NS3 displays increased helicase activity, suggesting that the protease domain plays an assisting role in the RNA unwinding process. The structural interaction between the helicase and protease domain has been assessed using small angle X-ray scattering on full length NS3, disclosing that the protease and helicase domains build a rather elongated molecular assembly differing from that observed in the NS3 protein from hepatitis C virus.
- Department of Biomolecular Sciences and Biotechnology, CNR-INFM, University of Milano, Via Celoria 26, 20133, Milano, Italy.
Organizational Affiliation: