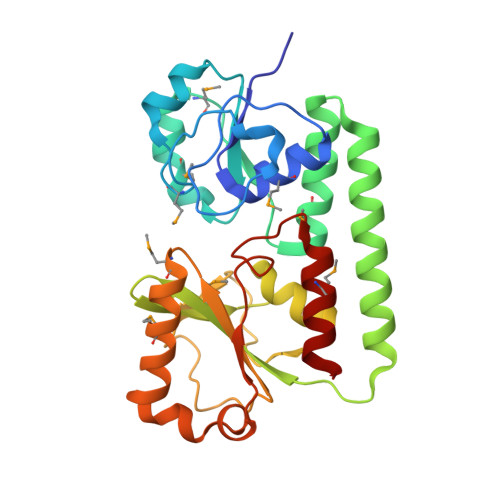
Heme Coordination by Staphylococcus aureus IsdE.
Grigg, J.C., Vermeiren, C.L., Heinrichs, D.E., Murphy, M.E.(2007) J Biological Chem 282: 28815-28822
- PubMed: 17666394 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M704602200
- Primary Citation Related Structures:
2Q8P, 2Q8Q - PubMed Abstract:
Staphylococcus aureus is a Gram-positive bacterial pathogen and a leading cause of hospital acquired infections. Because the free iron concentration in the human body is too low to support growth, S. aureus must acquire iron from host sources. Heme iron is the most prevalent iron reservoir in the human body and a predominant source of iron for S. aureus. The iron-regulated surface determinant (Isd) system removes heme from host heme proteins and transfers it to IsdE, the cognate substrate-binding lipoprotein of an ATP-binding cassette transporter, for import and subsequent degradation. Herein, we report the crystal structure of the soluble portion of the IsdE lipoprotein in complex with heme. The structure reveals a bi-lobed topology formed by an N- and C-terminal domain bridged by a single alpha-helix. The structure places IsdE as a member of the helical backbone metal receptor superfamily. A six-coordinate heme molecule is bound in the groove established at the domain interface, and the heme iron is coordinated in a novel fashion for heme transporters by Met(78) and His(229). Both heme propionate groups are secured by H-bonds to IsdE main chain and side chain groups. Of these residues, His(229) is essential for IsdE-mediated heme uptake by S. aureus when growth on heme as a sole iron source is measured. Multiple sequence alignments of homologues from several other Gram-positive bacteria, including the human pathogens pyogenes, Bacillus anthracis, and Listeria monocytogenes, suggest that these other systems function equivalently to S. aureus IsdE with respect to heme binding and transport.
- Department of Microbiology and Immunology, Life Sciences Institute, The University of British Columbia, Vancouver, British Columbia, Canada V6T 1Z3.
Organizational Affiliation: