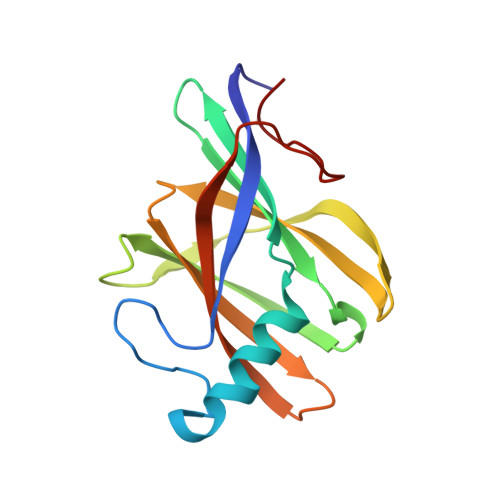
Bis-methionyl coordination in the crystal structure of the heme-binding domain of the streptococcal cell surface protein Shp.
Aranda IV, R., Worley, C.E., Liu, M., Bitto, E., Cates, M.S., Olson, J.S., Lei, B., Phillips Jr., G.N.(2007) J Mol Biology 374: 374-383
- PubMed: 17920629 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2007.08.058
- Primary Citation Related Structures:
2Q7A - PubMed Abstract:
Surface proteins Shr, Shp, and the ATP-binding cassette (ABC) transporter HtsABC are believed to make up the machinery for heme uptake in Streptococcus pyogenes. Shp transfers its heme to HtsA, the lipoprotein component of HtsABC, providing the only experimentally demonstrated example of direct heme transfer from a surface protein to an ABC transporter in Gram-positive bacteria. To understand the structural basis of heme transfer in this system, the heme-binding domain of Shp (Shp(180)) was crystallized, and its structure determined to a resolution of 2.1 A. Shp(180) exhibits an immunoglobulin-like beta-sandwich fold that has been recently found in other pathogenic bacterial cell surface heme-binding proteins, suggesting that the mechanisms of heme acquisition are conserved. Shp shows minimal amino acid sequence identity to these heme-binding proteins and the structure of Shp(180) reveals a unique heme-iron coordination with the axial ligands being two methionine residues from the same Shp molecule. A negative electrostatic surface of protein structure surrounding the heme pocket may serve as a docking interface for heme transfer from the more basic outer cell wall heme receptor protein Shr. The crystal structure of Shp(180) reveals two exogenous, weakly bound hemins, which form a large interface between the two Shp(180) molecules in the asymmetric unit. These "extra" hemins form a stacked pair with a structure similar to that observed previously for free hemin dimers in aqueous solution. The propionates of the protein-bound heme coordinate to the iron atoms of the exogenous hemin dimer, contributing to the stability of the protein interface. Gel filtration and analytical ultracentrifugation studies indicate that both full-length Shp and Shp(180) are monomeric in dilute aqueous solution.
- Department of Biomolecular Chemistry, University of Wisconsin-Madison, Madison, WI 53706, USA.
Organizational Affiliation: