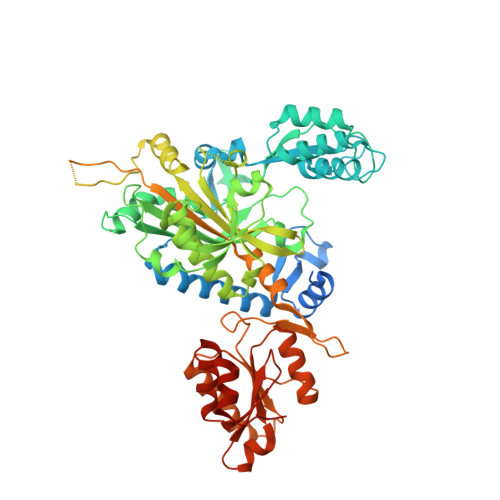
Crystal structure of human wildtype and S581L-mutant glycyl-tRNA synthetase, an enzyme underlying distal spinal muscular atrophy.
Cader, M.Z., Ren, J., James, P.A., Bird, L.E., Talbot, K., Stammers, D.K.(2007) FEBS Lett 581: 2959-2964
- PubMed: 17544401 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2007.05.046
- Primary Citation Related Structures:
2Q5H, 2Q5I - PubMed Abstract:
Dominant mutations in the ubiquitous enzyme glycyl-tRNA synthetase (GlyRS), including S581L, lead to motor nerve degeneration. We have determined crystal structures of wildtype and S581L-mutant human GlyRS. The S581L mutation is approximately 50A from the active site, and yet gives reduced aminoacylation activity. The overall structures of wildtype and S581L-GlyRS, including the active site, are very similar. However, residues 567-575 of the anticodon-binding domain shift position and in turn could indirectly affect glycine binding via the tRNA or alternatively inhibit conformational changes. Reduced enzyme activity may underlie neuronal degeneration, although a dominant-negative effect is more likely in this autosomal dominant disorder.
- Henry Wellcome Building for Gene Function, MRC Functional Genetics Unit, University of Oxford, South Parks Road, Oxford OX1 3QX, United Kingdom.
Organizational Affiliation: