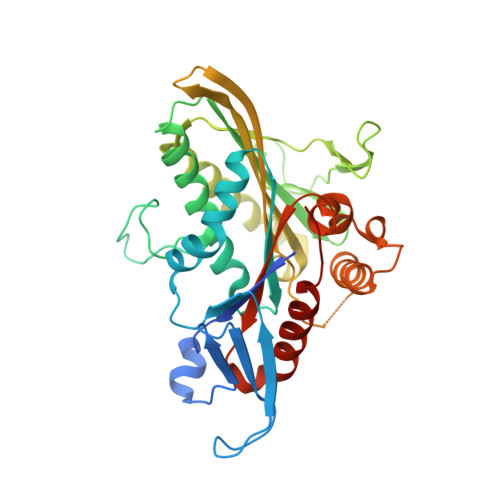
Kinesin spindle protein (KSP) inhibitors. Part 8: Design and synthesis of 1,4-diaryl-4,5-dihydropyrazoles as potent inhibitors of the mitotic kinesin KSP.
Roecker, A.J., Coleman, P.J., Mercer, S.P., Schreier, J.D., Buser, C.A., Walsh, E.S., Hamilton, K., Lobell, R.B., Tao, W., Diehl, R.E., South, V.J., Davide, J.P., Kohl, N.E., Yan, Y., Kuo, L.C., Li, C., Fernandez-Metzler, C., Mahan, E.A., Prueksaritanont, T., Hartman, G.D.(2007) Bioorg Med Chem Lett 17: 5677-5682
- PubMed: 17766111 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2007.07.074
- Primary Citation Related Structures:
2Q2Y, 2Q2Z - PubMed Abstract:
Inspired by previous efforts in the pyrazolobenzoxazine class of KSP inhibitors, the design and synthesis of 1,4-diaryl-4,5-dihydropyrazole inhibitors of KSP are described. Crystallographic evidence of binding mode and in vivo potency data is also highlighted.
- Department of Medicinal Chemistry, Merck Research Laboratories, PO Box 4, Sumneytown Pike, West Point, PA 19486, USA. anthony_roecker@merck.com
Organizational Affiliation: