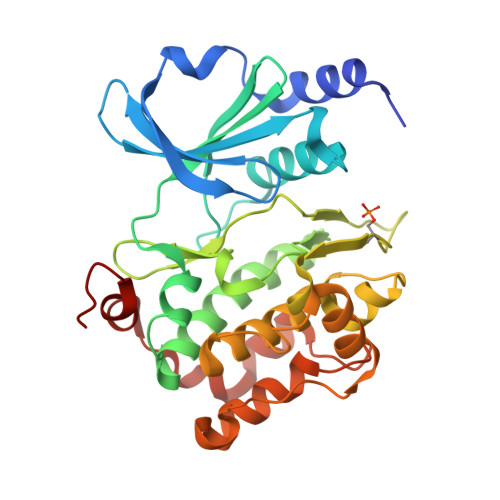
Structure of human p21 activating kinase 4 (PAK4) in complex with a consensus peptide.
Filippakopoulos, P., Eswaran, J., Turnbull, A., Papagrigoriou, E., Pike, A.W., von Delft, F., Sundstrom, M., Edwards, A., Arrowsmith, C.H., Weigelt, J., Knapp, S.To be published.