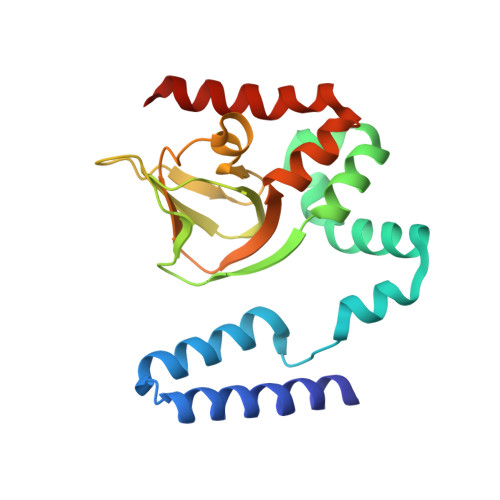
Structure and rearrangements in the carboxy-terminal region of SpIH channels.
Flynn, G.E., Black, K.D., Islas, L.D., Sankaran, B., Zagotta, W.N.(2007) Structure 15: 671-682
- PubMed: 17562314 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2007.04.008
- Primary Citation Related Structures:
2PTM, 2Q0A - PubMed Abstract:
Hyperpolarization-activated cyclic nucleotide-modulated (HCN) ion channels regulate the spontaneous firing activity and electrical excitability of many cardiac and neuronal cells. The modulation of HCN channel opening by the direct binding of cAMP underlies many physiological processes such as the autonomic regulation of the heart rate. Here we use a combination of X-ray crystallography and electrophysiology to study the allosteric mechanism for cAMP modulation of HCN channels. SpIH is an invertebrate HCN channel that is activated fully by cAMP, but only partially by cGMP. We exploited the partial agonist action of cGMP on SpIH to reveal the molecular mechanism for cGMP specificity of many cyclic nucleotide-regulated enzymes. Our results also elaborate a mechanism for the allosteric conformational change in the cyclic nucleotide-binding domain and a mechanism for partial agonist action. These mechanisms will likely extend to other cyclic nucleotide-regulated channels and enzymes as well.
- Department of Physiology and Biophysics, Howard Hughes Medical Institute, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: