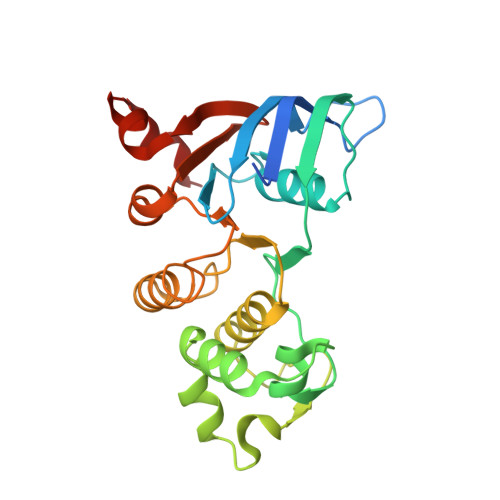
Structures of a minimal human CFTR first nucleotide-binding domain as a monomer, head-to-tail homodimer, and pathogenic mutant.
Atwell, S., Brouillette, C.G., Conners, K., Emtage, S., Gheyi, T., Guggino, W.B., Hendle, J., Hunt, J.F., Lewis, H.A., Lu, F., Protasevich, I.I., Rodgers, L.A., Romero, R., Wasserman, S.R., Weber, P.C., Wetmore, D., Zhang, F.F., Zhao, X.(2010) Protein Eng Des Sel 23: 375-384
- PubMed: 20150177 Search on PubMed
- DOI: https://doi.org/10.1093/protein/gzq004
- Primary Citation Related Structures:
2PZE, 2PZF, 2PZG - PubMed Abstract:
Upon removal of the regulatory insert (RI), the first nucleotide binding domain (NBD1) of human cystic fibrosis transmembrane conductance regulator (CFTR) can be heterologously expressed and purified in a form that remains stable without solubilizing mutations, stabilizing agents or the regulatory extension (RE). This protein, NBD1 387-646(Delta405-436), crystallizes as a homodimer with a head-to-tail association equivalent to the active conformation observed for NBDs from symmetric ATP transporters. The 1.7-A resolution X-ray structure shows how ATP occupies the signature LSGGQ half-site in CFTR NBD1. The DeltaF508 version of this protein also crystallizes as a homodimer and differs from the wild-type structure only in the vicinity of the disease-causing F508 deletion. A slightly longer construct crystallizes as a monomer. Comparisons of the homodimer structure with this and previously published monomeric structures show that the main effect of ATP binding at the signature site is to order the residues immediately preceding the signature sequence, residues 542-547, in a conformation compatible with nucleotide binding. These residues likely interact with a transmembrane domain intracellular loop in the full-length CFTR channel. The experiments described here show that removing the RI from NBD1 converts it into a well-behaved protein amenable to biophysical studies yielding deeper insights into CFTR function.
- Eli Lilly & Co., 10300 Campus Point Drive, San Diego, CA 92121, USA.
Organizational Affiliation: