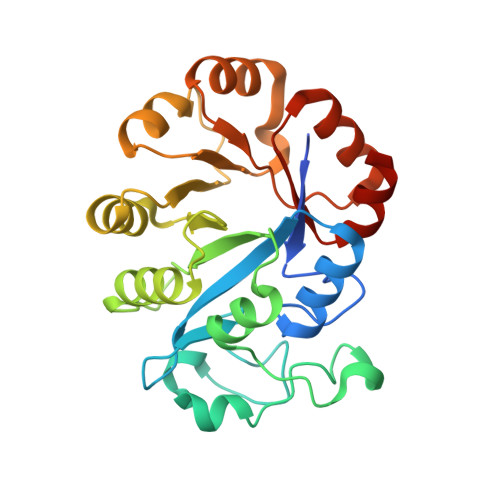
Crystal structure of glycerophosphodiester phosphodiesterase (GDPD) from Thermoanaerobacter tengcongensis, a metal ion-dependent enzyme: insight into the catalytic mechanism.
Shi, L., Liu, J.F., An, X.M., Liang, D.C.(2008) Proteins 72: 280-288
- PubMed: 18214974 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21921
- Primary Citation Related Structures:
2PZ0 - PubMed Abstract:
Glycerophosphodiester phosphodiesterase (GDPD; EC 3.1.4.46) catalyzes the hydrolysis of a glycerophosphodiester to an alcohol and glycerol 3-phosphate in glycerol metabolism. It has an important role in the synthesis of a variety of products that participate in many biochemical pathways. We report the crystal structure of the Thermoanaerobacter tengcongensis GDPD (ttGDPD) at 1.91 A resolution, with a calcium ion and glycerol as a substrate mimic coordinated at this calcium ion (PDB entry 2pz0). The ttGDPD dimer with an intermolecular disulfide bridge and two hydrogen bonds is considered as the potential functional unit. We used site-directed mutagenesis to characterize ttGDPD as a metal ion-dependent enzyme, identified a cluster of residues involved in substrate binding and the catalytic reaction, and we propose a possible general acid-base catalytic mechanism for ttGDPD. Superposing the active site with the homologous structure GDPD from Agrobacterium tumefaciens (PDB entry 1zcc), which binds a sulfate ion in the active site, the sulfate ion can represent the phosphate moiety of the substrate, simulating the binding mode of the true substrate of GDPD.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, People's Republic of China.
Organizational Affiliation: