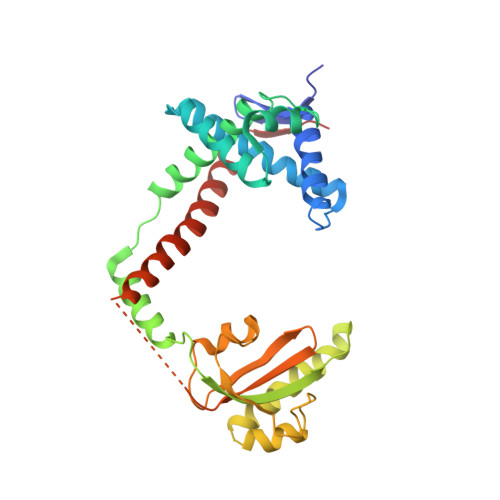
The Periplasmic Bacterial Molecular Chaperone SurA Adapts its Structure to Bind Peptides in Different Conformations to Assert a Sequence Preference for Aromatic Residues.
Xu, X., Wang, S., Hu, Y.X., McKay, D.B.(2007) J Mol Biology 373: 367-381
- PubMed: 17825319 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2007.07.069
- Primary Citation Related Structures:
2PV1, 2PV2, 2PV3 - PubMed Abstract:
The periplasmic molecular chaperone protein SurA facilitates correct folding and maturation of outer membrane proteins in Gram-negative bacteria. It preferentially binds peptides that have a high fraction of aromatic amino acids. Phage display selections, isothermal titration calorimetry and crystallographic structure determination have been used to elucidate the basis of the binding specificity. The peptide recognition is imparted by the first peptidyl-prolyl isomerase (PPIase) domain of SurA. Crystal structures of complexes between peptides of sequence WEYIPNV and NFTLKFWDIFRK with the first PPIase domain of the Escherichia coli SurA protein at 1.3 A resolution, and of a complex between the dodecapeptide and a SurA fragment lacking the second PPIase domain at 3.4 A resolution, have been solved. SurA binds as a monomer to the heptapeptide in an extended conformation. It binds as a dimer to the dodecapeptide in an alpha-helical conformation, predicated on a substantial structural rearrangement of the SurA protein. In both cases, side-chains of aromatic residues of the peptides contribute a large fraction of the binding interactions. SurA therefore asserts a recognition preference for aromatic amino acids in a variety of sequence configurations by adopting alternative tertiary and quaternary structures to bind peptides in different conformations.
- Department of Structural Biology, Stanford University School of Medicine, Stanford, CA 94305, USA.
Organizational Affiliation: