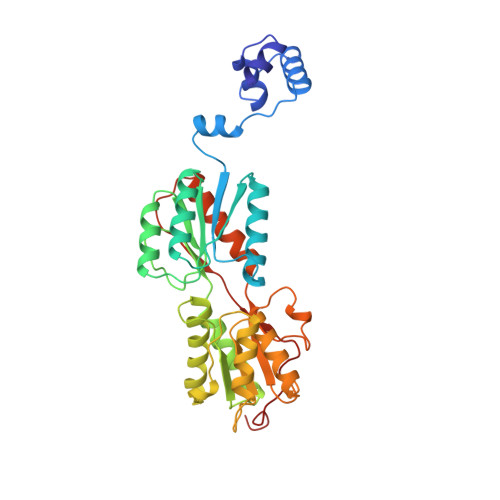
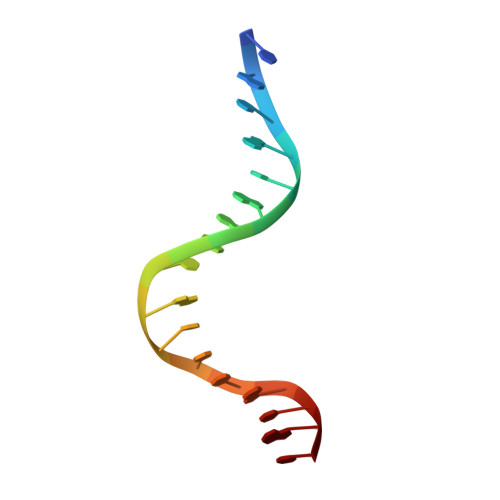
Crystal structure of LacI member, PurR, bound to DNA: minor groove binding by alpha helices.
Schumacher, M.A., Choi, K.Y., Zalkin, H., Brennan, R.G.(1994) Science 266: 763-770
- PubMed: 7973627 Search on PubMed
- DOI: https://doi.org/10.1126/science.7973627
- Primary Citation Related Structures:
1PNR, 2PUA, 2PUB, 2PUC, 2PUD - PubMed Abstract:
The three-dimensional structure of a ternary complex of the purine repressor, PurR, bound to both its corepressor, hypoxanthine, and the 16-base pair purF operator site has been solved at 2.7 A resolution by x-ray crystallography. The bipartite structure of PurR consists of an amino-terminal DNA-binding domain and a larger carboxyl-terminal corepressor binding and dimerization domain that is similar to that of the bacterial periplasmic binding proteins. The DNA-binding domain contains a helix-turn-helix motif that makes base-specific contacts in the major groove of the DNA. Base contacts are also made by residues of symmetry-related alpha helices, the "hinge" helices, which bind deeply in the minor groove. Critical to hinge helix-minor groove binding is the intercalation of the side chains of Leu54 and its symmetry-related mate, Leu54', into the central CpG-base pair step. These residues thereby act as "leucine levers" to pry open the minor groove and kink the purF operator by 45 degrees.
- Department of Biochemistry and Molecular Biology, Oregon Health Sciences University, Portland 97201-3098.
Organizational Affiliation: