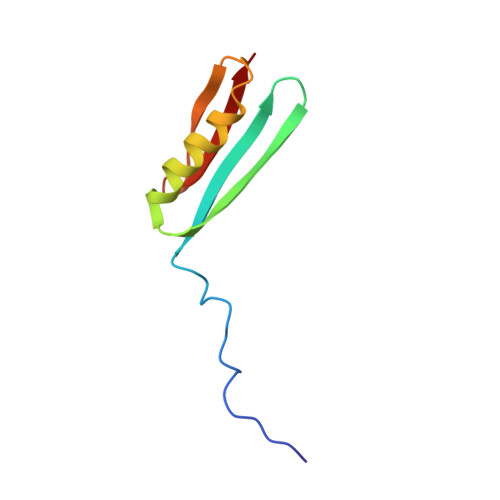
Three-dimensional solution structure of an immunoglobulin light chain-binding domain of protein L. Comparison with the IgG-binding domains of protein G.
Wikstrom, M., Drakenberg, T., Forsen, S., Sjobring, U., Bjorck, L.(1994) Biochemistry 33: 14011-14017
- PubMed: 7947810 Search on PubMed
- DOI: https://doi.org/10.1021/bi00251a008
- Primary Citation Related Structures:
2PTL - PubMed Abstract:
Protein L is a multidomain protein expressed at the surface of some strains of the anaerobic bacterial species Peptostreptococcus magnus. It has affinity for immunoglobulin (Ig) through interaction with framework structures in the variable Ig light chain domain. The Ig-binding activity is located to five homologous repeats called B1-B5 in the N-terminal part of the protein. We have determined the three-dimensional solution structure of the 76 amino acid residue long B1 domain using NMR spectroscopy and distance geometry-restrained simulated annealing. The domain is composed of a 15 amino acid residue long disordered N-terminus followed by a folded portion comprising an alpha-helix packed against a four-stranded beta-sheet. These secondary structural elements are well determined with a backbone atomic root mean square deviation from their mean of 0.54 A. The B domains of protein L show very limited sequence homology to the domains of streptococcal protein G interacting with the heavy chains of IgG. However, despite this fact, and their different binding properties, the fold of the B1 domain was found to be similar to the fold of the IgG-binding protein G domains [Wikström, M., Sjöbring, U., Kastern, W., Björck, L., Drakenberg, T., & Forsén, S. (1993) Biochemistry 32, 3381-3386]. In the present study, the solution structure of the B1 domain enabled a more detailed comparison which can explain the different Ig-binding specificities of these two bacterial surface proteins. Among the differences observed, the alpha-helix orientation is the most striking.(ABSTRACT TRUNCATED AT 250 WORDS)
- Department of Physical Chemistry 2, Lund University, Sweden.
Organizational Affiliation: