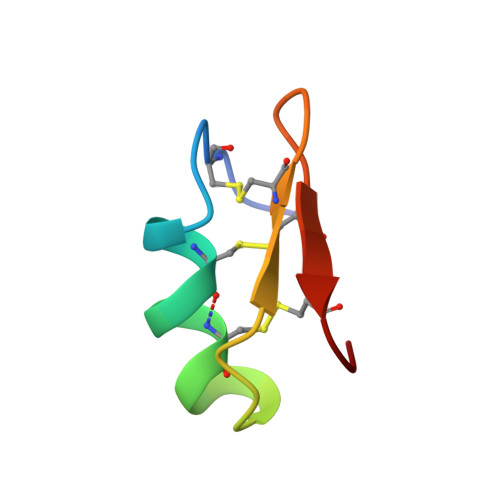
Solution structure for Pandinus toxin K-alpha (PiTX-K alpha), a selective blocker of A-type potassium channels.
Tenenholz, T.C., Rogowski, R.S., Collins, J.H., Blaustein, M.P., Weber, D.J.(1997) Biochemistry 36: 2763-2771
- PubMed: 9062103 Search on PubMed
- DOI: https://doi.org/10.1021/bi9628432
- Primary Citation Related Structures:
2PTA - PubMed Abstract:
PiTX-K alpha, a 35-residue peptide recently isolated from the venom of Pandinus imperator, blocks the rapidly inactivating (A-type) K+ channel(s) in rat brain synaptosomes and the cloned Kv 1.2 potassium channel at very low toxin concentrations (6 nM and 32 pM, respectively) [Rogowski, R. S., Collins, J. H., O'Neil, T. J., Gustafson, T. A., Werkman, T. A., Rogawski, M. A., Tenenholz, T. C., Weber, D. J., & Blaustein, M. P. (1996) Mol. Pharmacol. 50, 1167-1177]. The three-dimensional structure of PiTX-K alpha was determined using NMR spectroscopy in order to understand its selectivity and affinity toward K+ channels. PiTX-K alpha was found to have an alpha-helix from residues 10 to 21 and two beta-strands (betaI, 26-28; betaII, 33-35) connected by a type II beta-turn to form a small antiparallel beta-sheet. Three disulfide bonds, which are conserved in all members of the charybdotoxin family (alpha-K toxins), anchor one face of the alpha-helix to the beta-sheet. The N-terminal portion of PiTX-K alpha has three fewer residues than other alpha-K toxins such as charybdotoxin. Rather than forming a third beta-strand as found for other alpha-K toxins, the N-terminal region of PiTX-K alpha adopts an extended conformation. This structural difference in PiTX-K alpha together with differences in sequence at Pro-10, Tyr-14, and Asn-25 (versus Ser-10, Trp-14, and Arg-25 in CTX) may explain why PiTX-K alpha does not block maxi-K+ channels. Differences in three-dimensional structure between PiTX-K alpha and charybdotoxin are also observed in both the tight turn and the loop that connects the first beta-strand to the alpha-helix. As a result, side chains of two residues (Tyr-23 and Arg-31) are in regions of PiTX-K alpha that probably interact with rapidly inactivating A-type K+ channels. The analogous residues in charybdotoxin are positioned differently on the toxin surface. Thus, the locations of Tyr-23 and Arg-31 side chains in PiTX-K alpha could explain why this toxin blocks A-type channels at much lower concentrations than does charybdotoxin.
- Department of Biochemistry, University of Maryland School of Medicine, Baltimore 21201, USA.
Organizational Affiliation: