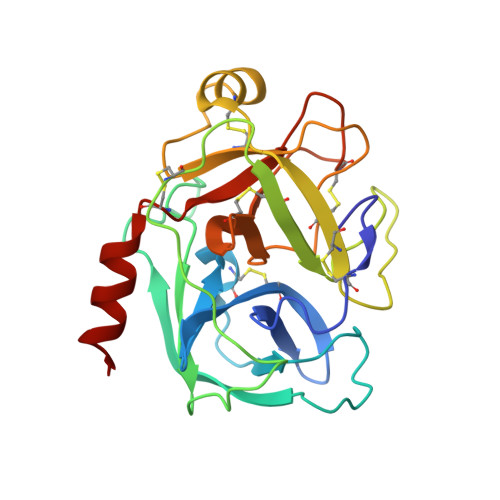
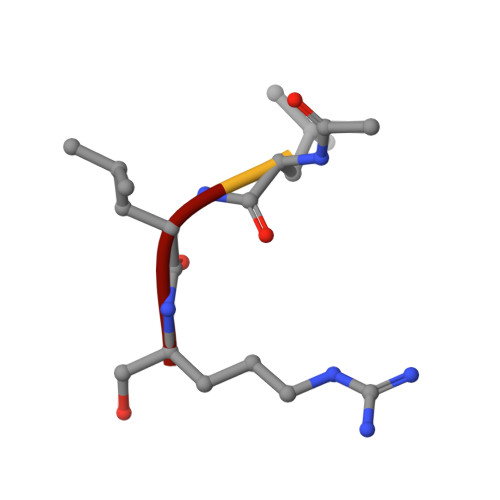
Structural basis of the zinc inhibition of human tissue kallikrein 5.
Debela, M., Goettig, P., Magdolen, V., Huber, R., Schechter, N.M., Bode, W.(2007) J Mol Biology 373: 1017-1031
- PubMed: 17881000
- DOI: https://doi.org/10.1016/j.jmb.2007.08.042
- Primary Citation Related Structures:
2PSX, 2PSY - PubMed Abstract:
Human kallikrein 5 (hK5) is a member of the tissue kallikrein family of serine peptidases. It has trypsin-like substrate specificity, is inhibited by metal ions, and is abundantly expressed in human skin, where it is believed to play a central role in desquamation. To further understand the interaction of hK5 with substrates and metal ions, active recombinant hK5 was crystallized in complex with the tripeptidyl aldehyde inhibitor leupeptin, and structures at 2.3 A resolution were obtained with and without Zn2+. While the overall structure and the specificity of S1 pocket for basic side-chains were similar to that of hK4, a closely related family member, both differed in their interaction with Zn2+. Unlike hK4, the 75-loop of hK5 is not structured to bind a Zn2+. Instead, Zn2+ binds adjacent to the active site, becoming coordinated by the imidazole rings of His99 and His96 not present in hK4. This zinc binding is accompanied by a large shift in the backbone conformation of the 99-loop and by large movements of both His side-chains. Modeling studies show that in the absence of bound leupeptin, Zn2+ is likely further coordinated by the imidazolyl side-chain of the catalytic His57 which can, similar to equivalent His57 imidazole groups in the related rat kallikrein proteinase tonin and in an engineered metal-binding rat trypsin, rotate out of its triad position to provide the third co-ordination site of the bound Zn2+, rendering Zn2+-bound hK5 inactive. In solution, this mode of binding likely occurs in the presence of free and substrate saturated hK5, as kinetic analyses of Zn2+ inhibition indicate a non-competitive mechanism. Supporting the His57 re-orientation, Zn2+ does not fully inhibit hK5 hydrolysis of tripeptidyl substrates containing a P2-His residue. The P2 and His57 imidazole groups would lie next to each other in the enzyme-substrate complex, indicating that incomplete inhibition is due to competition between both imidazole groups for Zn2+. The His96-99-57 triad is thus suggested to be responsible for the Zn2+-mediated inhibition of hK5 catalysis.
- Max-Planck-Institut für Biochemie, Proteinase Research Group, Am Klopferspitz 18, 82152 Martinsried, Germany.
Organizational Affiliation: