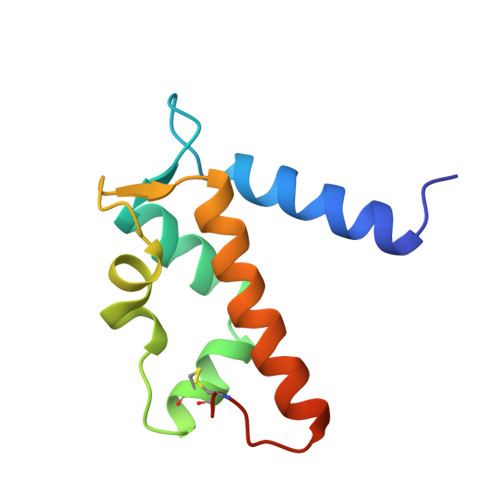
Zinc-binding site of an S100 protein revealed. Two crystal structures of Ca2+-bound human psoriasin (S100A7) in the Zn2+-loaded and Zn2+-free states.
Brodersen, D.E., Nyborg, J., Kjeldgaard, M.(1999) Biochemistry 38: 1695-1704
- PubMed: 10026247 Search on PubMed
- DOI: https://doi.org/10.1021/bi982483d
- Primary Citation Related Structures:
2PSR, 3PSR - PubMed Abstract:
The crystal structure of human psoriasin (S100A7) in the native, calcium-bound form has been determined from two crystal forms of the protein crystallized with and without divalent zinc. The overall structures of the dimeric protein closely resemble the previously determined holmium-substituted structure. The structures also reveal a zinc-binding site of the protein, which is formed by three histidines and an aspartate residue. Together, these residues coordinate the zinc ion in a way similar to the pattern seen in certain metalloproteases and in particular the collagenase family of proteins. Sequence comparison suggests that this zinc site is present in a number of the remaining members of the S100 family. The structure of S100A7 crystallized in the absence of zinc further shows that loss of zinc results in a reorganization of the adjacent empty and distorted EF-hand loop, causing it to resemble a calcium-loaded EF-hand.
- Macromolecular Crystallography, Institute of Molecular and Structural Biology, Aarhus University, Denmark.
Organizational Affiliation: