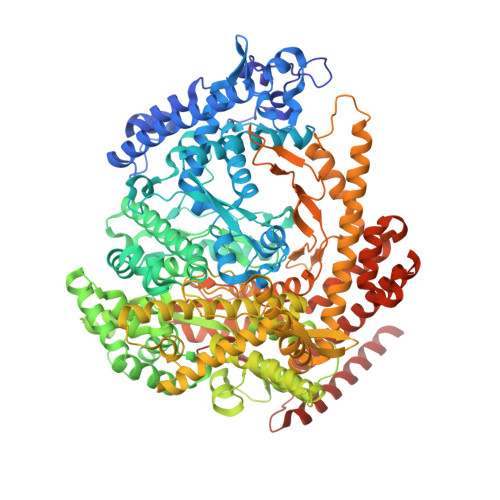
X-ray crystal structure of polymerase domain of the bacteriophage N4 virion RNA polymerase
Murakami, K.S., Davydova, E.K., Rothman-Denes, L.B.(2008) Proc Natl Acad Sci U S A 15: 5046-5051
- PubMed: 18362338 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0712325105
- Primary Citation Related Structures:
2PO4 - PubMed Abstract:
Coliphage N4 virion RNA polymerase (vRNAP), which is injected into the host upon infection, transcribes the phage early genes from promoters that have a 5-bp stem-3 nt loop hairpin structure. Here, we describe the 2.0-A resolution x-ray crystal structure of N4 mini-vRNAP, a member of the T7-like, single-unit RNAP family and the minimal component having all RNAP functions of the full-length vRNAP. The structure resembles a "fisted right hand" with Fingers, Palm and Thumb subdomains connected to an N-terminal domain. We established that the specificity loop extending from the Fingers along with W129 of the N-terminal domain play critical roles in hairpin-promoter recognition. A comparison with the structure of the T7 RNAP initiation complex reveals that the pathway of the DNA to the active site is blocked in the apo-form vRNAP, indicating that vRNAP must undergo a large-scale conformational change upon promoter DNA binding and explaining the highly restricted promoter specificity of vRNAP that is essential for phage early transcription.
- Department of Biochemistry and Molecular Biology, Pennsylvania State University, University Park, PA 16802, USA. kum14@psu.edu
Organizational Affiliation: